 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | SMil_00008897-RA_Salv | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Lamiaceae; Nepetoideae; Mentheae; Salvia
|
||||||||
| Family | MIKC_MADS | ||||||||
| Protein Properties | Length: 272aa MW: 30687.3 Da PI: 8.4772 | ||||||||
| Description | MIKC_MADS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SRF-TF | 84.3 | 7.3e-27 | 9 | 58 | 1 | 50 |
S---SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-TTSEEEEEE CS
SRF-TF 1 krienksnrqvtfskRrngilKKAeELSvLCdaevaviifsstgklyeys 50
k+ien rqvtfskRr+g++KKA+EL vLC+aevaviifs+tgkl+e++
SMil_00008897-RA_Salv 9 KKIENVNSRQVTFSKRRAGLFKKANELAVLCEAEVAVIIFSNTGKLFEFA 58
68***********************************************8 PP
| |||||||
| 2 | K-box | 61.6 | 3e-21 | 104 | 185 | 19 | 100 |
K-box 19 qelakLkkeienLqreqRhllGedLesLslkeLqqLeqqLekslkkiRskKnellleqieelqkkekelqeenkaLrkklee 100
+e++ Lk+eie+L+ +q +l+G+dL+ +s +eL+ Le+qL ++l i+++K++lll+++ + +++e+++ en++Lr+++ee
SMil_00008897-RA_Salv 104 KEADILKEEIEKLKLKQLRLMGKDLTGMSSQELHLLERQLSEGLMCIKERKEQLLLQELAQCRMQEQRAVLENETLRRQVEE 185
6999***************************************************************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50066 | 31.553 | 1 | 61 | IPR002100 | Transcription factor, MADS-box |
| SMART | SM00432 | 1.1E-39 | 1 | 60 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00265 | 7.28E-41 | 2 | 72 | No hit | No description |
| SuperFamily | SSF55455 | 2.09E-32 | 2 | 77 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 2.0E-27 | 3 | 23 | IPR002100 | Transcription factor, MADS-box |
| PROSITE pattern | PS00350 | 0 | 3 | 57 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF00319 | 1.4E-25 | 10 | 57 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 2.0E-27 | 23 | 38 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 2.0E-27 | 38 | 59 | IPR002100 | Transcription factor, MADS-box |
| PROSITE profile | PS51297 | 11.333 | 99 | 189 | IPR002487 | Transcription factor, K-box |
| Pfam | PF01486 | 7.2E-16 | 104 | 183 | IPR002487 | Transcription factor, K-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010262 | Biological Process | somatic embryogenesis | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048577 | Biological Process | negative regulation of short-day photoperiodism, flowering | ||||
| GO:0060862 | Biological Process | negative regulation of floral organ abscission | ||||
| GO:0060867 | Biological Process | fruit abscission | ||||
| GO:0071365 | Biological Process | cellular response to auxin stimulus | ||||
| GO:2000692 | Biological Process | negative regulation of seed maturation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005737 | Cellular Component | cytoplasm | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 272 aa Download sequence Send to blast |
MGRGKIEIKK IENVNSRQVT FSKRRAGLFK KANELAVLCE AEVAVIIFSN TGKLFEFANS 60 SMKKILSRYK KCVETTKSSA AELEPEELCS KFFTLSLQKP PQPKEADILK EEIEKLKLKQ 120 LRLMGKDLTG MSSQELHLLE RQLSEGLMCI KERKEQLLLQ ELAQCRMQEQ RAVLENETLR 180 RQVEELRGFF PPTSSPTPPL CIEYHSSVIK SDPAMKEGSR SPETACNGGL AEEDSDTTLQ 240 LGLPYGNGSR KRKMIDGETQ SSTSETQFRL HG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5f28_A | 5e-20 | 1 | 69 | 1 | 69 | MEF2C |
| 5f28_B | 5e-20 | 1 | 69 | 1 | 69 | MEF2C |
| 5f28_C | 5e-20 | 1 | 69 | 1 | 69 | MEF2C |
| 5f28_D | 5e-20 | 1 | 69 | 1 | 69 | MEF2C |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 248 | 253 | SRKRKM |
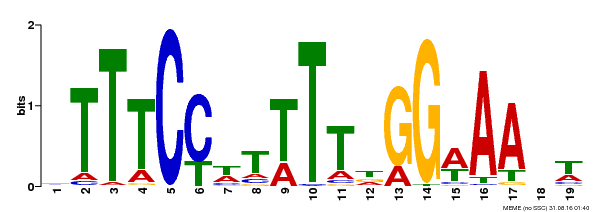
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00508 | DAP | Transfer from AT5G13790 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011091774.1 | 1e-126 | agamous-like MADS-box protein AGL15 | ||||
| TrEMBL | A0A4D8Z0G3 | 1e-125 | A0A4D8Z0G3_SALSN; MADS-box transcription factor, plant | ||||
| STRING | XP_006482951.1 | 3e-96 | (Citrus sinensis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA8775 | 22 | 26 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G13790.1 | 1e-53 | AGAMOUS-like 15 | ||||



