 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Potri.002G188700.1 | ||||||||
| Common Name | POPTR_0002s18970g | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Populus
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 975aa MW: 108029 Da PI: 6.1992 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 132.7 | 1.2e-41 | 150 | 227 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+C +dls+ak+yhrrhkvCe+hska+++lv +++qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+++
Potri.002G188700.1 150 VCQVEDCGVDLSNAKDYHRRHKVCEMHSKASKALVGNVMQRFCQQCSRFHVLQEFDEGKRSCRRRLAGHNKRRRKTNP 227
6**************************************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 2.5E-33 | 146 | 212 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.308 | 148 | 225 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 5.49E-39 | 149 | 230 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 4.5E-30 | 151 | 224 | IPR004333 | Transcription factor, SBP-box |
| Gene3D | G3DSA:1.25.40.20 | 5.9E-6 | 746 | 857 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 2.33E-7 | 750 | 858 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 2.37E-7 | 751 | 858 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016021 | Cellular Component | integral component of membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 975 aa Download sequence Send to blast |
MEARFGGEPH HFYAMGPTDM RAVGKRGLEW DLNDWKWDGD LFIASPLNPV PSTSVSRPFF 60 PLGVGTGVPA TGNSSNSSSS CSDEVNLGVE KGKRELEKRR RVVVIDDDNL NDQETGGLSL 120 KLGGQRDVGN WEGSSGKKTK LVGGGLSRAV CQVEDCGVDL SNAKDYHRRH KVCEMHSKAS 180 KALVGNVMQR FCQQCSRFHV LQEFDEGKRS CRRRLAGHNK RRRKTNPDTV GNGSSMNDDQ 240 NSGYLLISLL RILSNMHSNR SDETTDQDLL THLLRSLASH SVEHGGRNMF GPLQEPRDLS 300 TSFGNSEQVM PVHDAYGANI QTTSSLKPSI PNNFAVYSEV RESTAGQVKM NNFDLNDICV 360 DSDDGTEDIE RSPAPVNART SSLDCPSWVQ QDSHQSSPPQ TSRNSDSASA QSPSSSSGEA 420 QSRTDRIVFK LFGKEPNDFP LVLRAQILDW LSHSPTDIES YIRPGCIILT IYLHQAEAAW 480 EELCCGLGSS LSRLLAVSED TFWRTGWIYI RVQHQIAFVY NGQVVVDTSL PLTSNNYSKI 540 LSVKPIAITA SERAEFLIKG VNLSRPATRL LCAVEGNYMV QENRQEVMDG VDSFKGHDEV 600 QCVNFSCSIP MVTGRGFIEI EDHGFSSSFF PFLVAEEDVC SEIRMLEGVL ETETDADFEE 660 TEKMEAKNQA MNFVHEMSWL LHRSQLKSRL GCSDPSMNLF PLRRFKWLME FSMDHEWCAV 720 VGKLLNILHN GIVGTEEHSS LNVALSEMGL LHRAVRRNSR SLVELLLRYV PEKFGSKDTA 780 LVGGSHESIL FRPDVTGPAG LTPLHIAAGK DGSEDVLDTL TEDPGMVGIE AWKNAVDSTG 840 FTPEDYARLR GHYTYIHLVQ RKINKRQAVG GHVVLDIPSN LSNSNINEKQ NEGLSSSFEI 900 GQTALRPTQG NCKLCSQKVV YGIASRSQLY RPAMLSMVAI AAVCVCVALL FKSCPEVLYV 960 FRPFRWEMLD YGTS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 3e-30 | 150 | 224 | 10 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 97 | 101 | KRRRV |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Pth.11723 | 0.0 | bud| stem | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Expressed constitutively during plant development. {ECO:0000269|PubMed:10524240}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Binds specifically to the 5'-GTAC-3' core sequence. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:16095614}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
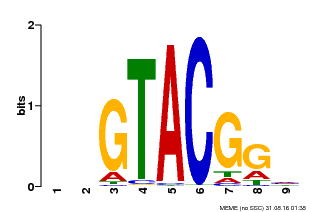
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Potri.002G188700.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024450080.1 | 0.0 | squamosa promoter-binding-like protein 1 isoform X2 | ||||
| Swissprot | Q9SMX9 | 0.0 | SPL1_ARATH; Squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | A0A2I4KKA4 | 0.0 | A0A2I4KKA4_POPTR; SBP family protein | ||||
| STRING | POPTR_0002s18970.1 | 0.0 | (Populus trichocarpa) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Potri.002G188700.1 |
| Entrez Gene | 7481436 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



