 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Peinf101Scf02665g01028.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Petunioideae; Petunia; Petunia integrifolia
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 1004aa MW: 110882 Da PI: 7.6128 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 129.5 | 1.2e-40 | 152 | 229 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+C+adls+ak+yhrrhkvC+vhska ++lv +++qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+++
Peinf101Scf02665g01028.1 152 VCQVEDCQADLSNAKDYHRRHKVCDVHSKAVSALVGNVMQRFCQQCSRFHILQEFDEGKRSCRRRLAGHNKRRRKTHP 229
6**************************************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 4.8E-32 | 146 | 214 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 31.949 | 150 | 227 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.96E-37 | 151 | 232 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 5.3E-29 | 153 | 226 | IPR004333 | Transcription factor, SBP-box |
| Gene3D | G3DSA:1.25.40.20 | 1.0E-6 | 775 | 895 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 5.12E-6 | 775 | 882 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 2.14E-6 | 776 | 883 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1004 aa Download sequence Send to blast |
METKIGGKIN QFYGPVVADL NLKAGGKKTM EWDLNDWNWD GDLFTAAPLN SVPSDCRSKQ 60 LFPIGSGVQE ITTGVSNSLS SGSDEMGLGI DKVRKELEKR RRAVVSEDDE LNDEAASLNL 120 KLGGQLYPIM ERGDGKSGKK SKVAGASSNR AVCQVEDCQA DLSNAKDYHR RHKVCDVHSK 180 AVSALVGNVM QRFCQQCSRF HILQEFDEGK RSCRRRLAGH NKRRRKTHPE NVANGASVND 240 ERGSNYLLIS LLRILANIHY QTKDQDLLSH LLRNLASLAS AANERNAAGL QSATPDLRNT 300 GTTSGEICLK DSPQPTGQCM SIPTPEVREK RMGMGGADCG IPQNRGASQP DSLYHRKESI 360 PANANVPATS LAGMKLSNID LNNSYDDSQD GIGKVHNPDA TMNLGNGSTG YPLWLCQEPH 420 KSSPTRISGN SGSTSTLSPS NSSGESRTDR IVFKLFGKDP SDFPTTLRKQ ILDWLAHSPT 480 DIESYIRPGC IILTIYLRMD KSIWEELYCD LSSSLTKLLN ASTDSFWRTG WVYARVQHRV 540 AFIFNGQVVL DTPLPVTSHR NCRISIIKPL AVSASEGVQF LVKGFNLSRP TTRLLCALEG 600 KYLVQGSCTD MMGGVDSYIE HDEPQSLGFP CIMPNITGRG FIEVEDHGLC SSFFPFIVAE 660 KDVCSEIRTL ENIIEVAETA DGSLGETEEL QARDQALDFI HEMGWLLHRS QLQLRLGSGS 720 NLNLFPFERF KWLIEFSVDH AWCAVVKNLL GIFFNGIVDV RQHSSLDIAL QEVGVLHRAV 780 RGNNRSVVEM LLRYSPSGVL DKSGVENQQG RGSYLFRPDT EGPGGLTPLH IVASLDGYDG 840 LLEALIDDPE QIGTEAWRSA RDSTGLTPND YASLRGHYSY IHIVQKKINQ KSSSGHVVVN 900 IPGTLLNSSF KQKLQDDHKS VKVGSLQMEK PLMKPIQRHC RQCERKFSYG NMGTTSVAIY 960 RPAMLSMVAI AAICVCVALL FKSSPEVLYV FKPFRWEQLK YGSS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 4e-31 | 144 | 226 | 2 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 221 | 227 | KRRRKTH |
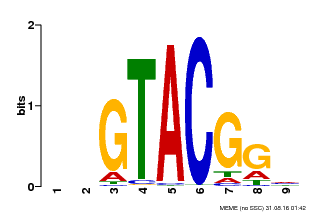
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HG975440 | 2e-56 | HG975440.1 Solanum pennellii chromosome ch01, complete genome. | |||
| GenBank | HG975441 | 2e-56 | HG975441.1 Solanum pennellii chromosome ch02, complete genome. | |||
| GenBank | HG975444 | 2e-56 | HG975444.1 Solanum pennellii chromosome ch05, complete genome. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_019253402.1 | 0.0 | PREDICTED: squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | A0A2K9ZXI9 | 0.0 | A0A2K9ZXI9_PETHY; Squamosa promoter binding-like protein 12d | ||||
| STRING | Solyc05g053240.2.1 | 0.0 | (Solanum lycopersicum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA1871 | 24 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||



