 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.9NG840500.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 686aa MW: 73768.7 Da PI: 6.5545 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 89.9 | 2.2e-28 | 199 | 252 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr++W+ eLH++Fv+av++L G++kA+Pk+ilelm+v+gLt+e+v+SHLQk+Rl
Pavir.9NG840500.1.p 199 KPRVVWSVELHQQFVNAVNHL-GIDKAVPKKILELMNVPGLTRENVASHLQKFRL 252
79*******************.********************************8 PP
| |||||||
| 2 | Response_reg | 77.6 | 4.5e-26 | 15 | 123 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTTSEEEEEESTTTHHHHHH CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepklpiivvtahgeeedale 88
vl+vdD+p+ + +l+++l + +y +v+++ +++ al++l+e++ +D+i+ D+ mp+mdG++ll++ e +lp+i+++a + + + +
Pavir.9NG840500.1.p 15 VLVVDDDPTCLVVLKRMLLECRY-DVTTCPQATRALTMLRENRrgFDVIISDVHMPDMDGFRLLEHVGLEM-DLPVIMMSADSRTDIVMK 102
89*********************.***************99999***********************6644.8***************** PP
HHHTTESEEEESS--HHHHHH CS
Response_reg 89 alkaGakdflsKpfdpeelvk 109
+k Ga d+l Kp+ +eel +
Pavir.9NG840500.1.p 103 GIKHGACDYLIKPVRMEELKN 123
******************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF036392 | 4.9E-180 | 1 | 668 | IPR017053 | Response regulator B-type, plant |
| Gene3D | G3DSA:3.40.50.2300 | 1.3E-42 | 12 | 148 | No hit | No description |
| SuperFamily | SSF52172 | 1.71E-36 | 12 | 137 | IPR011006 | CheY-like superfamily |
| SMART | SM00448 | 1.8E-30 | 13 | 125 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 43.037 | 14 | 129 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 3.7E-23 | 15 | 123 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 4.99E-27 | 16 | 128 | No hit | No description |
| PROSITE profile | PS51294 | 11.674 | 196 | 255 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 9.14E-20 | 196 | 256 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 1.1E-29 | 196 | 257 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 3.0E-25 | 199 | 252 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 7.8E-7 | 201 | 250 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0071368 | Biological Process | cellular response to cytokinin stimulus | ||||
| GO:0080113 | Biological Process | regulation of seed growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 686 aa Download sequence Send to blast |
MAEARGGDFP VGMKVLVVDD DPTCLVVLKR MLLECRYDVT TCPQATRALT MLRENRRGFD 60 VIISDVHMPD MDGFRLLEHV GLEMDLPVIM MSADSRTDIV MKGIKHGACD YLIKPVRMEE 120 LKNIWQHVVR KKFNENKDHE HSGSLDDTDR NRPTNNDNEY ASSANDGGDG SWKSQKKKRE 180 KEEDEADLEN GDPSSTSKKP RVVWSVELHQ QFVNAVNHLG IDKAVPKKIL ELMNVPGLTR 240 ENVASHLQKF RLYLKRIAQH HAGIPHPFVA PASSTKVAPL GGLEFQALAA SGQIPPQALA 300 ALQDELLGRP TSSLALPGRD QSSLQLAAIK GNKPHGEREI AFGQPIYKCP NNTYGAFPQS 360 SPGVGGLPSF GAWPNNKLGM TDSPSVLGNV NNSQNSNLLL HELQQQPDSL LSGTLHNIDV 420 KPSGIVMPSQ SLNAFPASEG ISPNQNPLVI PSQSQSFLAS ISPSMKHEPV LASSSPSNSL 480 LGGIDMVNQA STSQPLISTH GTNLPGLLNR SSNAVPSPGI SNFQSGNNPY VVNQNAMGVS 540 SRPPGVLKTE STDSLSRSYG YIGGSTSVDS GLLSQSKNPQ YGLLPNPNDV TGSWSPSQDI 600 DSCGNNLGQG YPGSTSSNYQ SSNVTLGKLP DQGRGRNHEF VGKGTCIPTR FAVDEVESPT 660 NNLSHNIGNS ADIVNPDMFG FSGQM* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 7e-24 | 195 | 258 | 1 | 64 | ARR10-B |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Pvr.878 | 0.0 | callus| leaf| root| stem | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specific DNA sequence. Functions as a response regulator involved in His-to-Asp phosphorelay signal transduction system. Phosphorylation of the Asp residue in the receiver domain activates the ability of the protein to promote the transcription of target genes. May directly activate some type-A response regulators in response to cytokinins. {ECO:0000250|UniProtKB:Q940D0}. | |||||
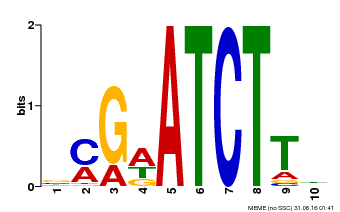
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00054 | PBM | Transfer from AT4G16110 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.9NG840500.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB060130 | 0.0 | AB060130.1 Zea mays ZmRR8 mRNA for response regulator 8, complete cds. | |||
| GenBank | EU975545 | 0.0 | EU975545.1 Zea mays clone 492147 two-component response regulator ARR1 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025794212.1 | 0.0 | two-component response regulator ORR21 | ||||
| Refseq | XP_025794213.1 | 0.0 | two-component response regulator ORR21 | ||||
| Swissprot | A2XE31 | 0.0 | ORR21_ORYSI; Two-component response regulator ORR21 | ||||
| TrEMBL | A0A3L6SBE0 | 0.0 | A0A3L6SBE0_PANMI; Two-component response regulator | ||||
| STRING | Pavir.Ia04946.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5796 | 37 | 54 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G16110.1 | 1e-115 | response regulator 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.9NG840500.1.p |



