 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pahal.I01317.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 970aa MW: 106090 Da PI: 5.5982 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 131.4 | 3.2e-41 | 151 | 228 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+CqvegC adl+ ak+yhrrhkvCe+h+ka++++v ++ qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+++
Pahal.I01317.1 151 ACQVEGCGADLTAAKDYHRRHKVCEMHAKASTAVVGNTVQRFCQQCSRFHLLQEFDEGKRSCRRRLAGHNRRRRKTRP 228
5**************************************************************************875 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 2.8E-34 | 144 | 213 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.624 | 149 | 226 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 7.19E-39 | 150 | 230 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 4.6E-30 | 152 | 225 | IPR004333 | Transcription factor, SBP-box |
| CDD | cd00204 | 1.62E-9 | 726 | 856 | No hit | No description |
| SMART | SM00248 | 0.29 | 751 | 780 | IPR002110 | Ankyrin repeat |
| Gene3D | G3DSA:1.25.40.20 | 4.8E-8 | 752 | 855 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 6.53E-8 | 753 | 857 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 710 | 797 | 827 | IPR002110 | Ankyrin repeat |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 970 aa Download sequence Send to blast |
MEAARLGAQS SHLYGSGLGE LDLNRRENRV FGWDLNDWNW DSERFVATPV PTAAANGSGL 60 NSSPSSSEEA EAEVARNGGH RGDTDKRKRV VVIDDDDRED QDMIVNGGRS LSLRIGGSAV 120 GVGVMESSDV NEDDRNGKKI RVQGGSSNGP ACQVEGCGAD LTAAKDYHRR HKVCEMHAKA 180 STAVVGNTVQ RFCQQCSRFH LLQEFDEGKR SCRRRLAGHN RRRRKTRPDI AIGGTASIED 240 KVSNYLLLSL IGICANLNSD SVQDSNSQEL LSTLLKNLGS VAKSLEPKEL CKLLEAYQSL 300 QNGSNAGTSG TANAAEEAAG PSNSKLPFVN GSHRGQASSS VVPVQSNATV VVTPEPASCK 360 LKDFDLNDTC NDMEGFEDGQ EGSPTPAFKA ADSPNCASWM QQDSTQSPPQ TSGNSDSTST 420 QSLSSSNGDA QCRTDKIVFK LFNKVPSDLP PVLRSQILGW LSSSPTDIES YIRPGCIILT 480 VYLRLVESAW RELSDNMSLH LDKLLNSSNG DFWASGLVFV IVRRQLAFMH NGQIMLDRPL 540 APSSHQYCKI LRVRPVAAPY SATINFRVEG FNLLSTSSRV ICSYEGRCIF QEDTDTVADD 600 AEYKDRDIEC LSFCCSIPGT RGRGFIEVED SGFSNGFFPF IIAEKDVCSE VSELESIFES 660 SSNEHAVVDD NARDQSLEFL NELGWLLHRV NRMSKQDETD TPLSAFNMWR FRKLGIFAME 720 REWCAVVEML LDFLFIGLVD VGSRSPEEMV LSENLLHAAV RTKSVKMVRF LLRYKPNKNS 780 KGTAQTYLFR PDALGPLTIT PLHIAAATSD AEDVLDALTD DPGLVGISAW SNARDETGFT 840 PEDYARQRGN DAYLNLVRKK IDKHVGKGHV VLGVPSSMCS VIPDGAKPGD VSLEICTPMS 900 ASVPRCLLCT QQARVYPNSG ARTFLYRPAM LTVMGVAVVC VCVGILLHTF PRVYAAPTFR 960 WELLERGPM* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 2e-31 | 145 | 225 | 4 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
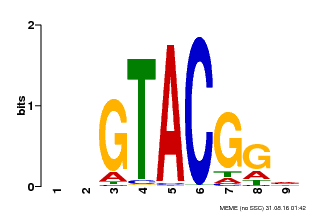
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT038852 | 0.0 | BT038852.1 Zea mays full-length cDNA clone ZM_BFb0336G12 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025793312.1 | 0.0 | squamosa promoter-binding-like protein 6 | ||||
| Swissprot | Q75LH6 | 0.0 | SPL6_ORYSJ; Squamosa promoter-binding-like protein 6 | ||||
| TrEMBL | A0A2S3IGN3 | 0.0 | A0A2S3IGN3_9POAL; Uncharacterized protein | ||||
| STRING | Pavir.J01500.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1998 | 38 | 92 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pahal.I01317.1 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



