 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PDK_30s1114731g003 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Coryphoideae; Phoeniceae; Phoenix
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 324aa MW: 34376.5 Da PI: 9.5961 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 70 | 3.8e-22 | 183 | 245 | 1 | 63 |
XXXXCHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 1 ekelkrerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkkevaklksev 63
e+elkrerrkq+NRe+ArrsR+RK+ae+e+L+ kv++L aeN+aL++e+++l++++ kl+ e+
PDK_30s1114731g003 183 ERELKRERRKQSNRESARRSRLRKQAETEQLAVKVETLGAENMALRSEISRLSENSNKLRLEN 245
89*********************************************************9887 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.20.5.170 | 6.0E-18 | 175 | 242 | No hit | No description |
| SMART | SM00338 | 2.5E-20 | 183 | 247 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF00170 | 1.9E-20 | 183 | 245 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 12.783 | 185 | 248 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 3.31E-10 | 186 | 241 | No hit | No description |
| CDD | cd14702 | 6.79E-20 | 188 | 236 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 190 | 205 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0005829 | Cellular Component | cytosol | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 324 aa Download sequence Send to blast |
PLLPPPFGTP YTAIYPAGGV YSHPSMPFGS HMQYQGIAPS PTISEAVVMA TPLSMEMPAK 60 SPGNKDKGPM KKLKGFDGHA TAMGNGSAEN RGGDGKVDAQ ANAAHGGETS ASSKMSLGIT 120 VAPPNISGKP VGTGPSSSPT PGMEFRVATT GKVQTSGTTI SPAAGAMMPG RNRGSSELWI 180 QDERELKRER RKQSNRESAR RSRLRKQAET EQLAVKVETL GAENMALRSE ISRLSENSNK 240 LRLENSTLME KLKTARLSQE EISTANMESE GAPSVVAENF LSMIDNSSSI SRSVQQEDEA 300 HEKSSGKLHQ LLDSNPRTDA VAAS |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 199 | 205 | RRSRLRK |
| 2 | 199 | 206 | RRSRLRKQ |
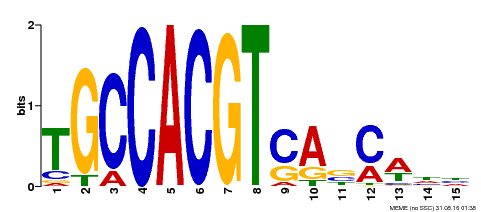
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00318 | DAP | Transfer from AT2G46270 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008798658.1 | 0.0 | common plant regulatory factor 1-like | ||||
| TrEMBL | A0A2H3YHM7 | 0.0 | A0A2H3YHM7_PHODC; common plant regulatory factor 1-like | ||||
| STRING | XP_008798658.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2776 | 38 | 88 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G46270.1 | 6e-60 | G-box binding factor 3 | ||||



