 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | ORGLA03G0387300.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 221aa MW: 23862.7 Da PI: 6.5094 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 58.8 | 1.3e-18 | 76 | 125 | 3 | 55 |
AP2 3 ykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
++GVr ++ +g+++AeIrd + ng r++lg+f++aeeAa a+++a+ +++g
ORGLA03G0387300.1 76 FRGVRKRP-WGKFAAEIRDSTRNG--VRVWLGTFDSAEEAALAYDQAAFAMRG 125
79******.**********44465..*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51032 | 22.682 | 75 | 133 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 2.4E-36 | 75 | 139 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 3.6E-30 | 75 | 134 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 2.22E-22 | 76 | 135 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 3.66E-29 | 76 | 133 | No hit | No description |
| Pfam | PF00847 | 1.7E-13 | 76 | 125 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.5E-10 | 76 | 87 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.5E-10 | 99 | 115 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0009507 | Cellular Component | chloroplast | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 221 aa Download sequence Send to blast |
MHCCMSLHPH RRHGDGDVDG SGSARLTAGL INFLESRRAG AMSTTNSSSS VSVPAMDAHG 60 QEEEEEPMQV QQQQAFRGVR KRPWGKFAAE IRDSTRNGVR VWLGTFDSAE EAALAYDQAA 120 FAMRGSAAVL NFPMEQVRRS MDMSLLQEGA SPVVALKRRH SMRAAAAGRR RKSAAPAPAD 180 QEGGGGVMEL EDLGPDYLEE LLAASQPIDI TCCTSPSHHS I |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5wx9_A | 4e-28 | 76 | 135 | 15 | 74 | Ethylene-responsive transcription factor ERF096 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 156 | 171 | KRRHSMRAAAAGRRRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the GCC-box cis-acting elements found in the promoter regions of ethylene-responsive genes (PubMed:23057995). Acts downstream of MYC2 in the jasmonate-mediated response to Botrytis cinerea infection (PubMed:28733419). With MYC2 forms a transcription module that regulates pathogen-responsive genes (PubMed:28733419). {ECO:0000269|PubMed:23057995, ECO:0000269|PubMed:28733419}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00286 | DAP | Transfer from AT2G31230 | Download |
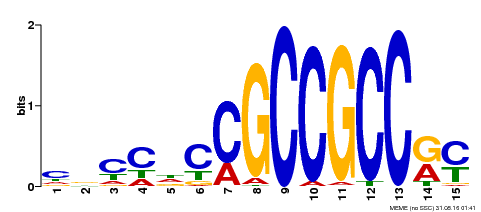
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | ORGLA03G0387300.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by wounding and infection with the fungal pathogen Botrytis cinerea. {ECO:0000269|PubMed:28733419}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC092263 | 0.0 | AC092263.7 Oryza sativa chromosome 3 BAC OSJNBa0033P04 genomic sequence, complete sequence. | |||
| GenBank | AK109390 | 0.0 | AK109390.2 Oryza sativa Japonica Group cDNA clone:006-211-G07, full insert sequence. | |||
| GenBank | AP014959 | 0.0 | AP014959.1 Oryza sativa Japonica Group DNA, chromosome 3, cultivar: Nipponbare, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015629379.1 | 1e-141 | ethylene-responsive transcription factor 15 | ||||
| Swissprot | A0A3Q7I5Y9 | 8e-50 | ERFC3_SOLLC; Ethylene-response factor C3 | ||||
| TrEMBL | I1PHM7 | 1e-161 | I1PHM7_ORYGL; Uncharacterized protein | ||||
| STRING | ORGLA03G0387300.1 | 1e-162 | (Oryza glaberrima) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP277 | 37 | 249 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G23240.1 | 1e-30 | ethylene response factor 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



