 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OMERI03G36740.2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 959aa MW: 104554 Da PI: 5.1719 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 130 | 8.4e-41 | 151 | 228 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+CqvegC+adl+ +++yhrrhkvCe+h+ka++++v ++ qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+++
OMERI03G36740.2 151 ACQVEGCTADLTGVRDYHRRHKVCEMHAKATTAVVGNTVQRFCQQCSRFHPLQEFDEGKRSCRRRLAGHNRRRRKTRP 228
5**************************************************************************875 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 6.3E-34 | 145 | 213 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.394 | 149 | 226 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.7E-38 | 150 | 230 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 1.6E-30 | 152 | 225 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF48403 | 2.02E-8 | 740 | 846 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 1.1E-7 | 743 | 849 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 9.20E-9 | 746 | 846 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016021 | Cellular Component | integral component of membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 959 aa Download sequence Send to blast |
MEAARVGAQS RHLYGGGLGE PDMDRRDKRL FGWDLNDWRW DSDRFVATPV PTAEASGLAL 60 NSSPSSSEEA GAALVRNVNA RGDSDKRKRV VVIDDDDVED DELVENGGGS LSLRIGGDAV 120 ANGAGVGGGA DEEDRNGKKI RVQGGSPSGP ACQVEGCTAD LTGVRDYHRR HKVCEMHAKA 180 TTAVVGNTVQ RFCQQCSRFH PLQEFDEGKR SCRRRLAGHN RRRRKTRPEV AVGGSAFTED 240 KDSTIVLAAD TAEHLRGQEL LSGLLRNLGA VAKSLDPKEL CKLLEACQSM QDGSNAGTSE 300 TANALVNTAV AEAAGPSNSK MPFVNGDQCG LASSSVVPVQ SKSPTVATPD PPACKFKDFD 360 LNDTCGGMEG FEDGYEGSPT PAFKTTDSPN CPSWMHQDST QSPPQTSGNS DSTSAQSLSS 420 SNGDAQCRTD KIVFKLFEKV PSDLPPVLRS QILGWLSSSP TDIESYIRPG CIILTVYLRL 480 VEPAWKELSD NMSSYLDKLL NSSTGNFWAS GLVFVMVRHQ IAFMHNGQLM LDRPLANSAH 540 HYCKILCVRP IAAPFSTKVN FRVEGLNLVS DSSRLICSFE GSCIFQEDTD NIVDDAEHDD 600 IEYLNFCCPL PSSRGRGFVE VEDGGFSNGF FPFIIAEQDI CSEVCELESI FESSSHEQAD 660 DDNARNQALE FLNELGWLLH RANIISKQNK VPLASFNIWR FRNLGIFAME REWCAVTKLL 720 LDFLFIGLVD IGSQSPEEVV LSENLLHAAV RMKSAQMVRF LLGYKPNESL KGTAETFLFR 780 PDAQGPSKFT PLHIAAATDD AEDVLDALTN DPGLVGINVW RNARDDAGFT PEDYARQRGN 840 DAYLNMVEKK INKHLGKGHV VLGVPSSIHP VITDGVKPGE VSLEIGMSVP PPAPRCNACS 900 RQALMYPNSM ARTFLYRPAM LTVMGIAVIC VCVGLLLHTC PKVYAAPTFR WELLERGPM |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 8e-31 | 145 | 225 | 4 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
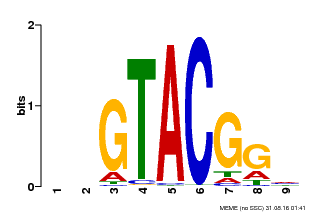
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK072164 | 0.0 | AK072164.1 Oryza sativa Japonica Group cDNA clone:J013130O11, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015631511.1 | 0.0 | squamosa promoter-binding-like protein 6 isoform X1 | ||||
| Swissprot | Q75LH6 | 0.0 | SPL6_ORYSJ; Squamosa promoter-binding-like protein 6 | ||||
| TrEMBL | A0A0E0D958 | 0.0 | A0A0E0D958_9ORYZ; Uncharacterized protein | ||||
| STRING | OGLUM03G39810.1 | 0.0 | (Oryza glumipatula) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



