 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OMERI01G06150.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | Dof | ||||||||
| Protein Properties | Length: 217aa MW: 22456.7 Da PI: 9.6212 | ||||||||
| Description | Dof family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-Dof | 121.3 | 3.4e-38 | 29 | 83 | 4 | 58 |
zf-Dof 4 kalkcprCdstntkfCyynnyslsqPryfCkaCrryWtkGGalrnvPvGggrrkn 58
+ l+cprC+stntkfCyynny+l+qPr+fCkaCrryWt+GGalrnvPvGgg+r++
OMERI01G06150.1 29 EGLACPRCESTNTKFCYYNNYNLAQPRHFCKACRRYWTRGGALRNVPVGGGTRNK 83
6789*************************************************86 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF02701 | 2.1E-32 | 31 | 83 | IPR003851 | Zinc finger, Dof-type |
| PROSITE profile | PS50884 | 28.416 | 31 | 85 | IPR003851 | Zinc finger, Dof-type |
| ProDom | PD007478 | 4.0E-23 | 33 | 81 | IPR003851 | Zinc finger, Dof-type |
| PROSITE pattern | PS01361 | 0 | 33 | 69 | IPR003851 | Zinc finger, Dof-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0042545 | Biological Process | cell wall modification | ||||
| GO:0045787 | Biological Process | positive regulation of cell cycle | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 217 aa Download sequence Send to blast |
MEVAEGRTVV AAVAAGGGGL GGGARTEAEG LACPRCESTN TKFCYYNNYN LAQPRHFCKA 60 CRRYWTRGGA LRNVPVGGGT RNKVAPAPCT GRRKRAAHAA APPPPTTTSV SSAPLPLMPP 120 AVAYELPFLP PPPPPPLTAM DPDRRLLDLG GSFTSLLAPA QLHNGHFTTG FLLGTMSSAP 180 PPPPPASTAT STPSPAPAAH PPVSDSIWAM GWPHLSI |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00402 | DAP | Transfer from AT3G50410 | Download |
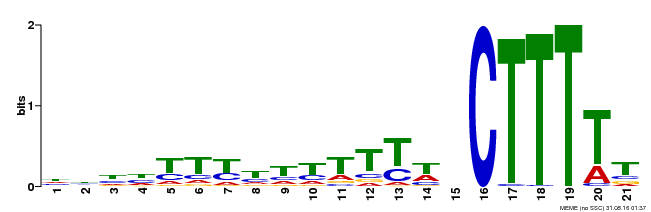
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP002537 | 0.0 | AP002537.2 Oryza sativa Japonica Group genomic DNA, chromosome 1, PAC clone:P0001B06. | |||
| GenBank | AP002746 | 0.0 | AP002746.2 Oryza sativa Japonica Group genomic DNA, chromosome 1, PAC clone:P0671B11. | |||
| GenBank | AP014957 | 0.0 | AP014957.1 Oryza sativa Japonica Group DNA, chromosome 1, cultivar: Nipponbare, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015645936.1 | 5e-90 | dof zinc finger protein DOF3.1 | ||||
| TrEMBL | A0A0E0BYI7 | 1e-152 | A0A0E0BYI7_9ORYZ; Uncharacterized protein | ||||
| STRING | OMERI01G06200.1 | 1e-153 | (Oryza meridionalis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP140 | 37 | 367 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G51700.1 | 2e-29 | DOF zinc finger protein 1 | ||||



