 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Niben101Scf00199g02008.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Nicotianoideae; Nicotianeae; Nicotiana
|
||||||||
| Family | MIKC_MADS | ||||||||
| Protein Properties | Length: 279aa MW: 31615.7 Da PI: 6.574 | ||||||||
| Description | MIKC_MADS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SRF-TF | 88.8 | 3e-28 | 9 | 59 | 1 | 51 |
S---SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-TTSEEEEEE- CS
SRF-TF 1 krienksnrqvtfskRrngilKKAeELSvLCdaevaviifsstgklyeyss 51
k ien nrqvtfskRr+g+lKKA+ELSvLCdaevaviifsst kl+e+ss
Niben101Scf00199g02008.1 9 KLIENVNNRQVTFSKRRAGLLKKANELSVLCDAEVAVIIFSSTSKLFEFSS 59
569**********************************************96 PP
| |||||||
| 2 | K-box | 57.8 | 4.8e-20 | 98 | 188 | 8 | 98 |
K-box 8 sleeakaeslqqelakLkkeienLqreqRhllGedLesLslkeLqqLeqqLekslkkiRskKnellleqieelqkkekelqeenk 92
+l+++ +++ q+e++ Lk+e+++L+ +q +llG+dL+ L l+eL+ Le+qL+++l i+++K+ ll++q+e+++++e+++ +e +
Niben101Scf00199g02008.1 98 QLQTHVQKQEQKEVDSLKDELSKLKIKQLRLLGKDLNGLGLNELRLLEHQLNEGLLAIKDRKEALLIQQLENSRRQEERAVSESE 182
3566667889*************************************************************************** PP
K-box 93 aLrkkl 98
+Lr++
Niben101Scf00199g02008.1 183 TLRRQA 188
***985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00432 | 2.3E-39 | 1 | 60 | IPR002100 | Transcription factor, MADS-box |
| PROSITE profile | PS50066 | 31.385 | 1 | 61 | IPR002100 | Transcription factor, MADS-box |
| SuperFamily | SSF55455 | 1.7E-32 | 2 | 75 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00265 | 1.14E-39 | 2 | 78 | No hit | No description |
| PRINTS | PR00404 | 9.3E-28 | 3 | 23 | IPR002100 | Transcription factor, MADS-box |
| PROSITE pattern | PS00350 | 0 | 3 | 57 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF00319 | 4.1E-25 | 11 | 57 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 9.3E-28 | 23 | 38 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 9.3E-28 | 38 | 59 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF01486 | 1.8E-14 | 100 | 187 | IPR002487 | Transcription factor, K-box |
| PROSITE profile | PS51297 | 11.418 | 104 | 194 | IPR002487 | Transcription factor, K-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010262 | Biological Process | somatic embryogenesis | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048577 | Biological Process | negative regulation of short-day photoperiodism, flowering | ||||
| GO:0060862 | Biological Process | negative regulation of floral organ abscission | ||||
| GO:0060867 | Biological Process | fruit abscission | ||||
| GO:0071365 | Biological Process | cellular response to auxin stimulus | ||||
| GO:2000692 | Biological Process | negative regulation of seed maturation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005737 | Cellular Component | cytoplasm | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 279 aa Download sequence Send to blast |
MGRGKIDIKL IENVNNRQVT FSKRRAGLLK KANELSVLCD AEVAVIIFSS TSKLFEFSST 60 SMKQTLSRYQ RCVASTEISA IERKSEDNQQ PQPQPQTQLQ THVQKQEQKE VDSLKDELSK 120 LKIKQLRLLG KDLNGLGLNE LRLLEHQLNE GLLAIKDRKE ALLIQQLENS RRQEERAVSE 180 SETLRRQARQ AFVEELRGLF PLSASLPPPY LEYNPLEKKY PIIKEGEESL DSDTACEDGV 240 DDEDSNTTLQ LGLPTIGRKR KKPEQESPSS NSENQVGSK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5f28_A | 5e-22 | 1 | 69 | 1 | 69 | MEF2C |
| 5f28_B | 5e-22 | 1 | 69 | 1 | 69 | MEF2C |
| 5f28_C | 5e-22 | 1 | 69 | 1 | 69 | MEF2C |
| 5f28_D | 5e-22 | 1 | 69 | 1 | 69 | MEF2C |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 257 | 261 | RKRKK |
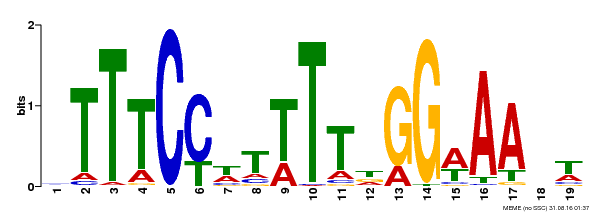
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00508 | DAP | Transfer from AT5G13790 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009787834.1 | 0.0 | PREDICTED: agamous-like MADS-box protein AGL15 | ||||
| Refseq | XP_016490205.1 | 0.0 | PREDICTED: agamous-like MADS-box protein AGL15 | ||||
| TrEMBL | A0A1S4BMX1 | 0.0 | A0A1S4BMX1_TOBAC; agamous-like MADS-box protein AGL15 | ||||
| TrEMBL | A0A1U7XLK3 | 0.0 | A0A1U7XLK3_NICSY; agamous-like MADS-box protein AGL15 | ||||
| STRING | XP_009787834.1 | 0.0 | (Nicotiana sylvestris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA8775 | 22 | 26 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G13790.1 | 4e-52 | AGAMOUS-like 15 | ||||



