 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Neem_6774_f_3 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Meliaceae; Azadirachta
|
||||||||
| Family | Nin-like | ||||||||
| Protein Properties | Length: 978aa MW: 107126 Da PI: 5.5913 | ||||||||
| Description | Nin-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | RWP-RK | 84.7 | 8.3e-27 | 560 | 620 | 2 | 52 |
RWP-RK 2 ekeisledlskyFslpikdAAkeLg..........vclTvLKriCRqyGIkRWPhRkiksl 52
ek+isle+l++yF++++kdAAk+Lg vc+T++KriCRq+GI+RWP+Rki+++
Neem_6774_f_3 560 EKSISLEVLQQYFAGSLKDAAKSLGdgsdkdscitVCPTTMKRICRQHGISRWPSRKINKV 620
799********************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51519 | 15.664 | 549 | 640 | IPR003035 | RWP-RK domain |
| Pfam | PF02042 | 6.2E-24 | 563 | 620 | IPR003035 | RWP-RK domain |
| SuperFamily | SSF54277 | 5.23E-23 | 875 | 968 | No hit | No description |
| SMART | SM00666 | 1.1E-25 | 882 | 964 | IPR000270 | PB1 domain |
| PROSITE profile | PS51745 | 26.192 | 882 | 964 | IPR000270 | PB1 domain |
| Gene3D | G3DSA:3.10.20.240 | 2.5E-26 | 883 | 963 | No hit | No description |
| Pfam | PF00564 | 3.7E-18 | 883 | 963 | IPR000270 | PB1 domain |
| CDD | cd06407 | 1.31E-31 | 883 | 963 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0010167 | Biological Process | response to nitrate | ||||
| GO:0042128 | Biological Process | nitrate assimilation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 978 aa Download sequence Send to blast |
MDLDVDLDLD NPWPLDQIGF VSNPMSPCLI SSSEQPSSPL WAFSDADNDD KIPGHVNYPL 60 FLKCNPTSET ENTKDNDKNR RFPSPLSGLI PLDSSDGYCM IKERMTQALR YFKDSTDQHV 120 LAQVWVPVKI GGRYVLTTSG QPFVLDPHSN GLHQYRMVSL MYMFSVDGEN DSELGLPGRV 180 FWQKLPEWTP NVQYYSSKEF SRLDHALNHN VRGTIALPVF EPSGQSCVGV IELIMTSQKI 240 NYAPEVDKVC KALEAVNLKS SEILDHPSTQ ICNGGRQNAL AEILEILSVV CETHKLPLAQ 300 TWVPCRHRSV LAYGGGFKKS CSSIDGSCMG KVCMSTTDVA FYVVDPHMWG FREACVEHHL 360 QKGQGVAGRS FFSLSSCFCK DITQFCKTEY PLVHYARMFG LTSCFAICLR SSHTGDDDYV 420 LEFFLPPVIV DSFEQQNLLG SMLATMKQHF QSLKVASGIE LEDDEVSIEI IEAFADKKIS 480 LRVGTIGIPQ SVQSPSEHNA LLNGQELLPL DLPEHQLMVN FEARNDGGNV VNVSGSNNPV 540 SFPENKNTRK SSERKRGKTE KSISLEVLQQ YFAGSLKDAA KSLGDGSDKD SCITVCPTTM 600 KRICRQHGIS RWPSRKINKV NRSLSKLKRV IESVQGADGT FGLTSLAASP LPVAVGSISW 660 PPVLNGSNQQ HSPSSKPSEP LGEKNGSPTY KTLGSDGHAG LDDRLSGGRM SDHKELIHEQ 720 NGLSPEIGKG TNSSKTGSGS REESAGSPTS HGSCQGSPAN ESTPAKDPLV TSIHEPCFKV 780 GGFPELVFQP TGELNLSAAF SIPDALVSTE PQEPFGGMLL EDAGSSKDLR NLCPAAADAI 840 ADERLPETSC TNPPCADVSP KQRLASLTQT MPHVHTRQEM KSMTIKATYR EDIIRFRISL 900 NSGIVELKEE VAKRLKLELG TFDIKYLDDD HEWVLIACDA DLQECLDISR SSGSSIIRLS 960 IHDIMPNLGS SCESTGEL |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in regulation of nitrate assimilation and in transduction of the nitrate signal. {ECO:0000269|PubMed:18826430}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00449 | DAP | Transfer from AT4G24020 | Download |
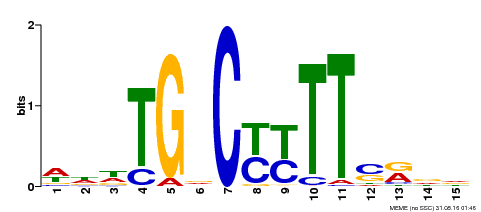
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Not regulated by the N source or by nitrate. {ECO:0000269|PubMed:18826430}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006439290.2 | 0.0 | LOW QUALITY PROTEIN: protein NLP7 | ||||
| Swissprot | Q84TH9 | 0.0 | NLP7_ARATH; Protein NLP7 | ||||
| TrEMBL | A0A173G8J1 | 0.0 | A0A173G8J1_PONTR; Transcription factor NLP7 | ||||
| STRING | XP_006476342.1 | 0.0 | (Citrus sinensis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2507 | 26 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G24020.1 | 0.0 | NIN like protein 7 | ||||



