 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Medtr7g028160.1 | ||||||||
| Common Name | MTR_7g028160 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Trifolieae; Medicago
|
||||||||
| Family | TCP | ||||||||
| Protein Properties | Length: 285aa MW: 30714.8 Da PI: 9.1195 | ||||||||
| Description | TCP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCP | 129.5 | 3.3e-40 | 44 | 153 | 2 | 107 |
TCP 2 agkkdrhskihTkvggRdRRvRlsaecaarfFdLqdeLGfdkdsktieWLlqqakpaikeltgtssssasec..eaesssssasnsssg..... 88
a+k+++++++hTkv+gR+RR+R++a+caar+F+L++eLG+++d++ti+WLlqq++p+i+++tgt++++as+ + ++ ++s+ +
Medtr7g028160.1 44 APKRTSNKDRHTKVEGRGRRIRMPALCAARIFQLTRELGHKSDGETIQWLLQQSEPSIIAATGTGTIPASALasSGNTLTPQGSSL--Ssglql 135
789******************************************************************77744333333333322..233443 PP
TCP 89 .....kaaksaakskksqksaasa 107
+ ++ ++a++
Medtr7g028160.1 136 ndrntW------AQTHQAHQAHQG 153
332221......222222222211 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03634 | 1.6E-33 | 48 | 151 | IPR005333 | Transcription factor, TCP |
| PROSITE profile | PS51369 | 28.171 | 50 | 104 | IPR017887 | Transcription factor TCP subgroup |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008361 | Biological Process | regulation of cell size | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0000987 | Molecular Function | core promoter proximal region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 285 aa Download sequence Send to blast |
MDPKNSKQQS QLSNMGENKE SETKNLQIVL SETTTKDETK KQLAPKRTSN KDRHTKVEGR 60 GRRIRMPALC AARIFQLTRE LGHKSDGETI QWLLQQSEPS IIAATGTGTI PASALASSGN 120 TLTPQGSSLS SGLQLNDRNT WAQTHQAHQA HQGHHVSSTS LWPHHHVGGF GFHQSSSSGG 180 LVATTVGENN SGNYFQKIGF SGFDMPTGTN LGVGGMSFTS ILGGANQQMP GLELGLSQDG 240 HIGVLNQQAL TQIYQQIGQN QTRVQHQNQQ NNNTTKDDSH SSEQ* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Mtr.11252 | 0.0 | leaf| root | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that binds to the site II motif (3'-TGGGCC/T-5') in the promoter of PCNA-2 and to 3'-GCCCG/A-5' elements in the promoters of cyclin CYCB1-1 and ribosomal protein genes. {ECO:0000269|PubMed:12631321, ECO:0000269|PubMed:16123132}. | |||||
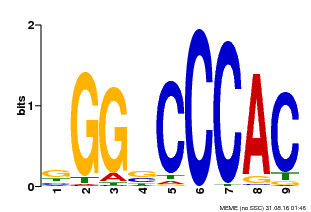
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00636 | PBM | Transfer from Glyma.19G095300 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Medtr7g028160.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC209532 | 0.0 | AC209532.2 Medicago truncatula chromosome 7 BAC clone mth2-62a7, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003622139.1 | 0.0 | transcription factor TCP20 | ||||
| Swissprot | Q9LSD5 | 3e-71 | TCP20_ARATH; Transcription factor TCP20 | ||||
| TrEMBL | G7L283 | 0.0 | G7L283_MEDTR; Putative transcription factor TCP family | ||||
| STRING | AES78357 | 0.0 | (Medicago truncatula) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3339 | 32 | 69 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G27010.1 | 9e-67 | TCP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Medtr7g028160.1 |
| Entrez Gene | 11439813 |



