 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Mapoly0001s0192.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Marchantiophyta; Marchantiopsida; Marchantiidae; Marchantiales; Marchantiaceae; Marchantia
|
||||||||
| Family | Dof | ||||||||
| Protein Properties | Length: 685aa MW: 68842.7 Da PI: 7.1677 | ||||||||
| Description | Dof family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-Dof | 119.3 | 1.4e-37 | 164 | 223 | 2 | 61 |
zf-Dof 2 kekalkcprCdstntkfCyynnyslsqPryfCkaCrryWtkGGalrnvPvGggrrknkks 61
+e+a++cprC+s +tkfCyynny+++qPr+fCk C+ryWt+GG+lrnvPvG+grrknk+
Mapoly0001s0192.1.p 164 PENAVACPRCHSFETKFCYYNNYNVNQPRHFCKHCQRYWTAGGTLRNVPVGAGRRKNKHG 223
67899****************************************************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| ProDom | PD007478 | 9.0E-26 | 162 | 221 | IPR003851 | Zinc finger, Dof-type |
| Pfam | PF02701 | 3.8E-31 | 166 | 222 | IPR003851 | Zinc finger, Dof-type |
| PROSITE profile | PS50884 | 28.053 | 168 | 222 | IPR003851 | Zinc finger, Dof-type |
| PROSITE pattern | PS01361 | 0 | 170 | 206 | IPR003851 | Zinc finger, Dof-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009658 | Biological Process | chloroplast organization | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048510 | Biological Process | regulation of timing of transition from vegetative to reproductive phase | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 685 aa Download sequence Send to blast |
MMDSPIKLFG KTIAKAAASA FTTSSGEADD AEAVPTTSSG GWVDLPSSSQ DVKENGAESS 60 RLGEINGGGV TFTRQEVKRG SSPDRTSSAC NSAEEEKLCA SPRGGGGATG GAGAELTPSA 120 SGGEAGAGDD EGSEEEDGSG MNGRSKKDGG DDGGKGGEKM PRRPENAVAC PRCHSFETKF 180 CYYNNYNVNQ PRHFCKHCQR YWTAGGTLRN VPVGAGRRKN KHGSTIAPRG QTPEVGTVRS 240 DAPDIASPLI ASKGGVGPVP VQVAVVQAAG SKGGIVRMAV PAASGDPSVL PFLRMQPSPM 300 DTASIFMASS DAGTSSDTVT VACKRGRLAT PGFSDASTGA NDMEQQQQQQ QQPSWGQQSS 360 TNDESACSPL TGIRSGGAQQ QQQQQQQQQQ QQQQQQHPAT NNGMAGPGGC WSAAGPQSSV 420 PPPGAAGAPY GFYNGGWPYG YNVGWSGPPA FCAGGVATAG TGMPGPPPPG NIAAWNGAGG 480 SVWTGVPWPM PGLAWGPPWA MAPWAMSAGH GGASMGSITQ TISMSSGAAA GMVTVPVPCS 540 TTASGGGPAS AAPPSSPSSS LGKHSRGGPP VGAPASSAAA AAACNGAGGA GGGGGGGGGG 600 RMEGGLWAPK SLRMADPEEA MRSSVWRILG VGSRADMTTA STFKAFQPRN ESKKISDDDQ 660 QAVQASLHAN PAAMSRSMNF HESN* |
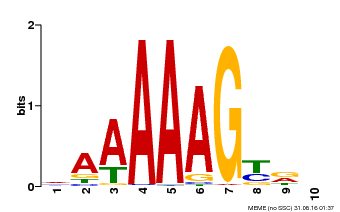
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00046 | PBM | Transfer from AT5G39660 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| TrEMBL | A0A2R6XVL7 | 0.0 | A0A2R6XVL7_MARPO; Uncharacterized protein | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP38 | 17 | 445 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G29160.1 | 2e-31 | Dof family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Mapoly0001s0192.1.p |



