 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Lj5g3v1548910.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Loteae; Lotus
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 1003aa MW: 111462 Da PI: 5.9401 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 125.9 | 1.6e-39 | 148 | 225 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+C adls+ak+yhrrhkvCe+hska+ +lv + +qrfCqqCsrfh+l+efDe+krsCrrrLa+hn+rrrk+++
Lj5g3v1548910.1 148 VCQVEDCCADLSRAKDYHRRHKVCEMHSKATRALVGNAMQRFCQQCSRFHMLQEFDEGKRSCRRRLAGHNKRRRKTNN 225
6**************************************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 1.4E-32 | 141 | 210 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 31.812 | 146 | 223 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.7E-36 | 147 | 227 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 8.4E-28 | 149 | 222 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF48403 | 1.2E-7 | 776 | 892 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.47E-6 | 780 | 891 | No hit | No description |
| Gene3D | G3DSA:1.25.40.20 | 3.7E-7 | 780 | 894 | IPR020683 | Ankyrin repeat-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1003 aa Download sequence Send to blast |
MEARFGAEAY HYYGVGASSD MRGMGKRSAE WDLNNWRWDG DLFIASRVNP VQEDGVGQQL 60 FPLVSGIPAS NSSSSCSEEV DLRDPMGSRE GERKRRVIVL EDDGLNEEGG TLSLKLGGHA 120 ADREVASWDG GNGKKSRVAG GGASNRAVCQ VEDCCADLSR AKDYHRRHKV CEMHSKATRA 180 LVGNAMQRFC QQCSRFHMLQ EFDEGKRSCR RRLAGHNKRR RKTNNEAVPN GNPLNDDQTS 240 SYLLISLLKI LSNMNSDRSD QTTDQDLLTH LLRSLASRNG EQGGNNLSNI LREPENLLRE 300 GGSSRESEMV PTLLSNGSQG SPTDIRQHQT VSMNKMQQEV VHAHDARATD QQLMSYIKPS 360 ISNTPPAYAE ARDSTAGQIK MNNFDLNDIY IDSDDGIEDL ERLPGTTNHV TSSLDYPWTQ 420 QDSHQSSPPQ TSRNSESGSA HSPSSSTGEA QCRTDRIVIK LFGKEPSDFP LILRAQILDW 480 LSHSPTDIES YIRPGCIVLT IYLRQAEAVW EEICYDLTSS LGRLLDVSDD TFWRTGWVHI 540 RVQHQMAFIF NGQVVIDTSL PFRSNNYSKI LTVSPIAAPA SKRVQFSVKG VNMIRPATRL 600 MCALEGKYLV CEDAHESMDQ YSEELDGLHC IQFSCAVPVT NGRGFIEIED QGLSSSFFPF 660 IVAEEDVCSD ICGLEPLLEL SETDPDIEGT GKIKAKSQAM DFIHEIGWLL HGSQLKSRMV 720 HLSSSTDLFP LKRFKWLMEF SMDHDWCAVV KKLLNLLLEG TVSTGDHPSL DLALSEMGLL 780 HRAVRRNSKQ LVELLLTYVP ENISNEVRPE GKALVEGEKQ SFLFRPDAAG PAGLTPLHIA 840 AGKDGSEDVL DALTNDPCMV GIEAWKNARD STGSTPEDYA RLRGHYTYIH LVQKKINKRQ 900 GAPHVVVEIP TNLTENDTNQ KQNESSTTFE IAKTRDQGHC KLCDIKLSCR TAVGRSLVYK 960 PAMLSMVAVA AVCVCVALLF KSSPEVLYMF RPFSWESLDF GTS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 5e-30 | 142 | 222 | 4 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Lja.25262 | 0.0 | pod| root | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Expressed constitutively during plant development. {ECO:0000269|PubMed:10524240}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Binds specifically to the 5'-GTAC-3' core sequence. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:16095614}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
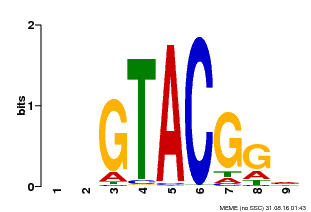
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Lj5g3v1548910.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001352074.1 | 0.0 | squamosa promoter-binding-like protein 1 | ||||
| Refseq | XP_028184385.1 | 0.0 | squamosa promoter-binding-like protein 1 | ||||
| Swissprot | Q9SMX9 | 0.0 | SPL1_ARATH; Squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | A0A0S3SQY1 | 0.0 | A0A0S3SQY1_PHAAN; Uncharacterized protein | ||||
| STRING | GLYMA02G01160.1 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1539 | 34 | 95 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



