 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LPERR02G18350.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Leersia
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 328aa MW: 34827.8 Da PI: 6.6922 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 54.7 | 2.5e-17 | 35 | 85 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkr.krfslgkfgtaeeAakaaiaarkkleg 55
++y+GVr+++ +gr++AeIrdp + r +lg+f+tae+Aa a++ a+++++g
LPERR02G18350.1 35 TRYRGVRRRP-WGRFAAEIRDPA----SkERRWLGTFDTAEQAACAYDVAARAMRG 85
68********.**********83....25999**********************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF00847 | 1.0E-10 | 35 | 85 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 1.79E-29 | 35 | 94 | No hit | No description |
| PROSITE profile | PS51032 | 23.182 | 36 | 93 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 8.5E-22 | 36 | 94 | IPR016177 | DNA-binding domain |
| SMART | SM00380 | 2.2E-35 | 36 | 99 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 1.8E-30 | 36 | 93 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.5E-9 | 37 | 48 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.5E-9 | 59 | 75 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009880 | Biological Process | embryonic pattern specification | ||||
| GO:0010084 | Biological Process | specification of organ axis polarity | ||||
| GO:0048825 | Biological Process | cotyledon development | ||||
| GO:0051726 | Biological Process | regulation of cell cycle | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005829 | Cellular Component | cytosol | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 328 aa Download sequence Send to blast |
MDDAAASAHA HQHRAKRPSP GGADGGAIVG SATGTRYRGV RRRPWGRFAA EIRDPASKER 60 RWLGTFDTAE QAACAYDVAA RAMRGTRART NFPVPSSAAV VSPAGGCWPW LTALPPSPHG 120 ATTATATQQQ PLNTFLLHNL LMSSSPHGCL LLHHAGHGHA HAHAHSHIRS HNPTRTPIST 180 PTPTPTPTPP VAITTPATTT TTTTAAGSAT FAAPGAYDDD DDEWGGLLRR EPPEAGLLQE 240 VLHGFYPANR PHASPVPAPK QVEMPYYEAS PWDVVEDCED GDVDDDGEYC GLPMMPQGLL 300 EDVIHCPPPY MEVLAPSSAI GRSRRGGT |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 2e-20 | 32 | 94 | 1 | 64 | ATERF1 |
| 3gcc_A | 2e-20 | 32 | 94 | 1 | 64 | ATERF1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00623 | PBM | Transfer from PK19363.1 | Download |
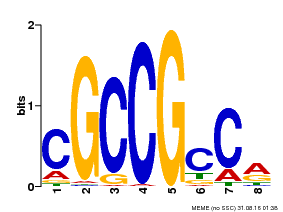
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LPERR02G18350.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP004023 | 5e-79 | AP004023.3 Oryza sativa Japonica Group genomic DNA, chromosome 2, BAC clone:OJ1126_D09. | |||
| GenBank | AP004079 | 5e-79 | AP004079.2 Oryza sativa Japonica Group genomic DNA, chromosome 2, BAC clone:OJ1067_B01. | |||
| GenBank | AP014958 | 5e-79 | AP014958.1 Oryza sativa Japonica Group DNA, chromosome 2, cultivar: Nipponbare, complete sequence. | |||
| GenBank | CP012610 | 5e-79 | CP012610.1 Oryza sativa Indica Group cultivar RP Bio-226 chromosome 2 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015624483.1 | 3e-79 | ethylene-responsive transcription factor ABI4 | ||||
| TrEMBL | A0A0D9VHS2 | 0.0 | A0A0D9VHS2_9ORYZ; Uncharacterized protein | ||||
| STRING | LPERR02G18350.1 | 0.0 | (Leersia perrieri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP7263 | 30 | 50 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G12980.1 | 6e-13 | ERF family protein | ||||



