 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KZV38026.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Gesneriaceae; Didymocarpoideae; Trichosporeae; Loxocarpinae; Dorcoceras
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 384aa MW: 41312 Da PI: 7.2953 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 68.7 | 9.2e-22 | 247 | 309 | 1 | 63 |
XXXXCHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 1 ekelkrerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkkevaklksev 63
e+elkrerrkq+NRe+ArrsR+RK+ae eeL+ kv++L++eN++ k+e+++l ++++klk e+
KZV38026.1 247 ERELKRERRKQSNRESARRSRLRKQAEHEELTIKVQALTSENMTIKSEINKLINNSEKLKLEN 309
89**********************************************************886 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF07777 | 3.8E-30 | 1 | 91 | IPR012900 | G-box binding protein, multifunctional mosaic region |
| Pfam | PF16596 | 4.9E-11 | 113 | 207 | No hit | No description |
| Gene3D | G3DSA:1.20.5.170 | 7.0E-16 | 239 | 306 | No hit | No description |
| Pfam | PF00170 | 1.6E-19 | 247 | 309 | IPR004827 | Basic-leucine zipper domain |
| SMART | SM00338 | 1.5E-18 | 247 | 311 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 12.231 | 249 | 312 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 3.13E-10 | 250 | 306 | No hit | No description |
| CDD | cd14702 | 1.86E-19 | 252 | 298 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 254 | 269 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0005829 | Cellular Component | cytosol | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 384 aa Download sequence Send to blast |
MGNSEEEKPC KPEALSSPAV DQNSIHVYPD WAAMQAYYGP RLAVPPYHNA VPASHAPPPY 60 MWGPPQSMIP PYGTPYAAFY AHGGVYAHPG VHIAGTTLSA ETPAKPPGNT DGGFVKKLKE 120 LDGLAMSIGN GNGNSTDDAA DNEVTQSEET EGASDGSNLS NGVTAGSGKK RSRTGTPNST 180 GNGKGKRLGQ DFEKVVNVTP AANMSAKTSE NTSTALMLKD PPDFDVKLSP TSVPQTLKPS 240 EAWLQNEREL KRERRKQSNR ESARRSRLRK QAEHEELTIK VQALTSENMT IKSEINKLIN 300 NSEKLKLENV ALMGKLKDAQ LGQTEDATLR KIGDFRPKPD GTVNLLARVN NNSGEGEAYE 360 NSSTGAKLHE LLDRSPRTDA VAAI |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 263 | 269 | RRSRLRK |
| 2 | 263 | 270 | RRSRLRKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds to the G-box-like motif (5'-ACGTGGC-3') of the chalcone synthase (CHS) gene promoter. G-box and G-box-like motifs are defined in promoters of certain plant genes which are regulated by such diverse stimuli as light-induction or hormone control. | |||||
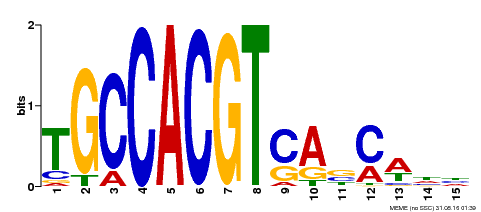
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00318 | DAP | Transfer from AT2G46270 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KZV38026.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By light. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011093357.1 | 1e-179 | common plant regulatory factor 1 isoform X3 | ||||
| Swissprot | Q99089 | 1e-120 | CPRF1_PETCR; Common plant regulatory factor 1 | ||||
| TrEMBL | A0A2Z7BUR2 | 0.0 | A0A2Z7BUR2_9LAMI; Transcriptional activator TAF-1 | ||||
| STRING | Migut.B01828.1.p | 1e-162 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA3113 | 24 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G46270.1 | 1e-104 | G-box binding factor 3 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



