 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KZV33652.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Gesneriaceae; Didymocarpoideae; Trichosporeae; Loxocarpinae; Dorcoceras
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 985aa MW: 109762 Da PI: 6.9406 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 130.5 | 5.9e-41 | 148 | 225 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+C+adls+ak+yhrrhkvC+vhska+++lv ++ qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+++
KZV33652.1 148 VCQVEDCKADLSNAKDYHRRHKVCDVHSKATKALVGNVWQRFCQQCSRFHVLEEFDEGKRSCRRRLAGHNKRRRKTHP 225
6**************************************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 1.1E-32 | 144 | 210 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.026 | 146 | 223 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.57E-37 | 147 | 228 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 2.7E-29 | 149 | 222 | IPR004333 | Transcription factor, SBP-box |
| Gene3D | G3DSA:1.25.40.20 | 9.9E-9 | 748 | 870 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 2.64E-10 | 748 | 871 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 3.32E-9 | 763 | 892 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 985 aa Download sequence Send to blast |
MEAKFGGKLH HFFGPVAPDL KAMGKKSVEW DLNDWKWDGD LFVAAPLNSV PSDCGNKQLF 60 PTGSETPAYN GASNSFSSGS DGIERERTEL EKRRRGEVET EQVNEECGSL NLKLGGQAYP 120 VGEVDVDRSE GKSGKKTKIS GVPSSRAVCQ VEDCKADLSN AKDYHRRHKV CDVHSKATKA 180 LVGNVWQRFC QQCSRFHVLE EFDEGKRSCR RRLAGHNKRR RKTHPENVVN AATLNDERGS 240 NYLLISLLKI LSNIHSNGSD QKDNQGLLSH LLRNLANLAG TNDERNPATL LPVSQDLPIV 300 DKYLENEKDP GRDVGQGVIA PASDLTPKRM LVGNSQGGIT DDASALTKKA NNSIKANAPD 360 APIERIRQFT IDLNNVYDDS QDCMDGLEDN VALENTRNVS PSSSFWLYKD SRNTQNSGNS 420 GSSSSQSPST SSGDEQSRTD RIVFKLFGKD PSDFPLVVRK QILDWLSNSP TEIESYIRPG 480 CIILTIYLRM DNAMWEELYC DISSSLRRLL DSSNDSFWKT GWIYSRVQNH ISFVYDGQVV 540 LDTPSHLKNH QGCRISSISP IAVCASESVQ FFVKGSNFSL VTSRLLCTID GKYLAQENCG 600 ARTGSAESFM EHDEIQSLNF SCTIPNIVGR GFIEVEDQGL SSSFFPFIVA EKDVCLEICT 660 LESIVGVTDG DTKKFEARNQ ALEFIHEMGW LLQRSRLKCR LGESSFNVGL FPLKRFRWLV 720 EFSIDHDWCA VVEKLLSIFF DGTVDSGKHT SILLALLDMG LLHQAVRRNC KSMVEFLLEY 780 RLSESFNKLG PKQNQAHDDD QYMFRPDSVG PGGLTPLHIA ACLDGRENVL DALTEDPRSV 840 GIDAWKNAKD STGLTPHDYA CFRGHYSYIH LVQRKLNKKL LNSHIVVDIP DNTVPAKQKI 900 GNTSKLGKSG VAFETERAQS CSECERKMGS YGNWRSSVRI YKPAMLSMVA IAAVCVCAAL 960 LFKTSPVVHP CFRPFRWELL KYGSE |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 6e-30 | 148 | 222 | 10 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 217 | 223 | KRRRKTH |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000269|PubMed:16554053}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
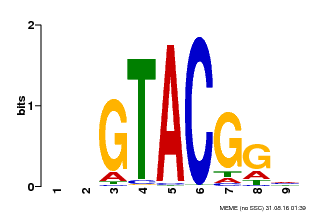
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KZV33652.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020547976.1 | 0.0 | LOW QUALITY PROTEIN: squamosa promoter-binding-like protein 1 | ||||
| Swissprot | Q9S7P5 | 0.0 | SPL12_ARATH; Squamosa promoter-binding-like protein 12 | ||||
| TrEMBL | A0A2Z7BH05 | 0.0 | A0A2Z7BH05_9LAMI; Squamosa promoter-binding-like protein 1-like | ||||
| STRING | Solyc05g053240.2.1 | 0.0 | (Solanum lycopersicum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA1871 | 24 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G60030.1 | 0.0 | squamosa promoter-binding protein-like 12 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



