 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Jcr4S02833.100 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Euphorbiaceae; Crotonoideae; Jatropheae; Jatropha
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 963aa MW: 106899 Da PI: 6.7072 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 132.5 | 1.4e-41 | 156 | 233 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+C +dls+ak+yhrrhkvCe+hska+++lv +++qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+++
Jcr4S02833.100 156 VCQVEDCGTDLSNAKDYHRRHKVCEMHSKASKALVGNVMQRFCQQCSRFHVLQEFDEGKRSCRRRLAGHNKRRRKTNP 233
6**************************************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 3.6E-33 | 151 | 218 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.214 | 154 | 231 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 8.89E-39 | 155 | 236 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 6.9E-30 | 157 | 230 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF48403 | 1.18E-7 | 753 | 862 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 1.7E-6 | 753 | 864 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.85E-7 | 755 | 862 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016021 | Cellular Component | integral component of membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 963 aa Download sequence Send to blast |
MEARFGGEAQ AHHFYSTGAT NLRAVGKRSL EWDLNDWKWD GDLFIANPLN PVPSGGMDRQ 60 FIPLATGISV NGNSSNSSSS CSDEVNLGIE KGKRELEKRR RVIVIEDDNL HGEEVGSLSL 120 KLGGHGYPVS EREMGNWEGN SGKKTKLVGG SMSRAVCQVE DCGTDLSNAK DYHRRHKVCE 180 MHSKASKALV GNVMQRFCQQ CSRFHVLQEF DEGKRSCRRR LAGHNKRRRK TNPDAVANGT 240 SLNDEQTSSY LLISLLRILS NMHSNRSDQV TDQDLLSHLL RSLASHTIDH GGRNISGLFQ 300 ESRDVLNDGT SFGNSEQVGH VHGANGATIQ TSSSIKPSIP NNYPAFSEVR DITGGQVKMN 360 NFDLNDIYID SDDGAEDIER SPVPTNMGTS SLDCPSWVQQ DSHQSSPPQT SGNSDSASAQ 420 SPSSSNGDAQ SRTDRIIFKL FGKEPNDFPL VLRAQILDWL SHSPTDIESY IRPGCVILTI 480 YLRQAETKWE ELCCNLSSSL SRLLDVSDDA FWRTGWVYIR VQHQIAFVYN GQVVVDTSLP 540 LRSSSYSRIL SVKPIAISAS ERAEFVIKGI NLSRPTTSIS GEDSLRCITS GIFLSNTTHP 600 WFWCNMLNLW TYTLVCALRW DYIEDQGFSS TFFPFIVAEE DFCSEIRMLE NVLDFTETNA 660 DVNGIGKMEA KNQAMDFIHE IGWLLHRSQL KYRLADLDPY TDLFPLKRFK WLMEFSVDHE 720 WCAVVKKLLN LLFNGVIGIG EHSSLNVALS EMGLLHRAVR KNSRSLVELL LRYVPEKSGA 780 VNNLLIGGSS ENFLFRPDVA GPAGLTPLHI AAGKDGSEDV LDALTDDTGM VGIEAWKNAR 840 DSTGFTPEDY ARLRGHYSYI HLVQKKINKK PAVGHVVLDI PGTLPDCSIN QKQNEGVSTS 900 FEIGQTAIRP IQRSCKLCHQ KLDYVTAGRS LLYRPAMLSM VAIAAVGIVG LRNKLRSSQP 960 DGL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 6e-33 | 150 | 230 | 4 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 97 | 101 | KRRRV |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000269|PubMed:16554053}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
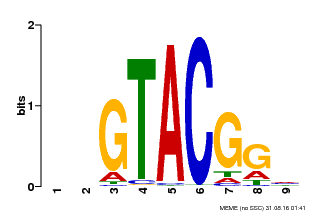
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012082979.1 | 0.0 | squamosa promoter-binding-like protein 1 | ||||
| Swissprot | Q9S7P5 | 0.0 | SPL12_ARATH; Squamosa promoter-binding-like protein 12 | ||||
| TrEMBL | A0A067K935 | 0.0 | A0A067K935_JATCU; Uncharacterized protein | ||||
| STRING | cassava4.1_000974m | 0.0 | (Manihot esculenta) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1539 | 34 | 95 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G60030.1 | 0.0 | squamosa promoter-binding protein-like 12 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



