 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Gorai.002G124400.1 | ||||||||
| Common Name | B456_002G124400, LOC105784599 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 946aa MW: 104093 Da PI: 5.1025 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 77.1 | 2e-24 | 149 | 250 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE- CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkld 85
f+k+lt sd++++g +++p++ ae+ ++++++ ++l+++d++ ++W++++iyr++++r++lt+GW+ Fv +++L++gD+v+F
Gorai.002G124400.1 149 FCKTLTASDTSTHGGFSVPRRAAEKLfpsldySMQPPT-QELVVRDLHDNTWTFRHIYRGQPKRHLLTTGWSLFVGSKRLRAGDSVLFI-- 236
99*********************999*****9555554.49************************************************.. PP
SSSEE..EEEEE-S CS
B3 86 grsefelvvkvfrk 99
++++ +l+v+v+r+
Gorai.002G124400.1 237 RDEKSQLLVGVRRA 250
4577778*****97 PP
| |||||||
| 2 | Auxin_resp | 114.5 | 9.4e-38 | 275 | 358 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalk.vkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
aahaa+++s+F+++YnPra++seFv+++ +++k+++ ++vsvGmRf m+fete+s +rr++Gt+vg+sdldp+rWp+SkWr+L+
Gorai.002G124400.1 275 AAHAAANRSPFTIFYNPRACPSEFVIPLPRYRKSVYgSQVSVGMRFGMMFETEESGKRRYMGTIVGISDLDPLRWPGSKWRNLQ 358
79**********************************99********************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 9.68E-47 | 136 | 278 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 4.0E-42 | 143 | 263 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 4.46E-22 | 148 | 249 | No hit | No description |
| SMART | SM01019 | 4.6E-24 | 149 | 251 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 13.183 | 149 | 251 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 7.0E-22 | 149 | 250 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 1.4E-32 | 275 | 358 | IPR010525 | Auxin response factor |
| PROSITE profile | PS51745 | 24.573 | 828 | 912 | IPR000270 | PB1 domain |
| SuperFamily | SSF54277 | 6.02E-9 | 830 | 907 | No hit | No description |
| Pfam | PF02309 | 5.9E-7 | 872 | 914 | IPR033389 | AUX/IAA domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0009908 | Biological Process | flower development | ||||
| GO:0009942 | Biological Process | longitudinal axis specification | ||||
| GO:0010305 | Biological Process | leaf vascular tissue pattern formation | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:0048507 | Biological Process | meristem development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016020 | Cellular Component | membrane | ||||
| GO:0042802 | Molecular Function | identical protein binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 946 aa Download sequence Send to blast |
MGSVVEEKIK QGGLVNVGAQ STLLEEMKLL KEMQDQSGTR KAINSELWHA CAGPLVSLPQ 60 VGSLVYYFPQ GHSEQVAVST KRMATSQIPN YPNLPSQLMC QVHNVTLHAD RDTDEIYAQM 120 SLQPVNSEKD VFPIPDFGLK LSKHPNEFFC KTLTASDTST HGGFSVPRRA AEKLFPSLDY 180 SMQPPTQELV VRDLHDNTWT FRHIYRGQPK RHLLTTGWSL FVGSKRLRAG DSVLFIRDEK 240 SQLLVGVRRA NRQQTTLPSS VLSADSMHIG VLAAAAHAAA NRSPFTIFYN PRACPSEFVI 300 PLPRYRKSVY GSQVSVGMRF GMMFETEESG KRRYMGTIVG ISDLDPLRWP GSKWRNLQVE 360 WDEPGCNDKQ NRVSAWEIET PESLFIFPSL TSSLKRPLYP GFSGAESEWG SLMKRPLLQF 420 PENGNGNLPY SMSNLCSEQL MKMMLKPQLV NHPGIFASPL HQIADVKVPP LEEMKNLQSK 480 SHTKPQVIQS ENMLIENRNL SHPVPDQPDP ITSNMSKINA NGNPHPANIL TQAGTGSSNE 540 KLKLESKHSA EQLTSTSECN EEKLVASTVN TTMSNQLSFP TQPHIPLQVQ NNPWSIQSQL 600 DSSVLQAHQM LVSQADISTL NSFLPFSDTD EWTSNLSSCQ PLSGAYKSPG PIPMVGLQDS 660 SAVFPVETDD SLTTVGEEIW DQKLNNCRVS SQADQLASFT QQDPCSLNSG GVRDLSDDSN 720 NQSGIYSSCL NIDVSNGCST VIDPFVSSAI LDEFCSLKDA DFQNPSDCLV GNFSSCQDVQ 780 SQITSASLAD SQAFSRQDLP DSSGGNIDFD DSGLLQNNSW KQTGPRVRTY TKVQKAGSVG 840 RSIDVTSFKN YDELISAIEC MFGLKGLLDD PRGSGWKLVY VDYENDVLLV GDDPWEEFVG 900 CVRCIRILSP TEVQQMSEEG MKLLNSAATV QGINGSNSEG SNANA* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldu_A | 0.0 | 3 | 383 | 6 | 392 | Auxin response factor 5 |
| Search in ModeBase | ||||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: In early embryo and during organ development. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in the whole plant with a lower expression in leaves. Detected in embryo axis, provascular tissues, procambium and some differentiated vascular regions of mature organs. {ECO:0000269|PubMed:10476078}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). Seems to act as transcriptional activator. Formation of heterodimers with Aux/IAA proteins may alter their ability to modulate early auxin response genes expression. Mediates embryo axis formation and vascular tissues differentiation. Functionally redundant with ARF7. May be necessary to counteract AMP1 activity. {ECO:0000269|PubMed:12036261, ECO:0000269|PubMed:14973283, ECO:0000269|PubMed:17553903}. | |||||
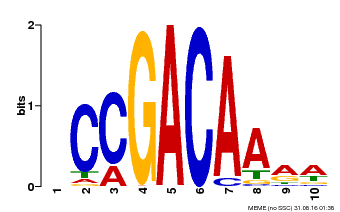
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00153 | DAP | Transfer from AT1G19850 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012465955.1 | 0.0 | PREDICTED: auxin response factor 5-like isoform X1 | ||||
| Swissprot | P93024 | 0.0 | ARFE_ARATH; Auxin response factor 5 | ||||
| TrEMBL | A0A0D2NM47 | 0.0 | A0A0D2NM47_GOSRA; Auxin response factor | ||||
| STRING | Gorai.002G124400.1 | 0.0 | (Gossypium raimondii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM5808 | 27 | 46 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G19850.1 | 0.0 | ARF family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Gorai.002G124400.1 |
| Entrez Gene | 105784599 |



