 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GSVIVT01027040001 | ||||||||
| Common Name | LOC100248302, VIT_15s0046g01440 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; rosids incertae sedis; Vitales; Vitaceae; Vitis
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 431aa MW: 45769.8 Da PI: 6.5279 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 65.8 | 7.5e-21 | 285 | 347 | 1 | 63 |
XXXXCHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 1 ekelkrerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkkevaklksev 63
e+e+krerrkq+NRe+ArrsR+RK+ae+eeL+ kv++L +eN+ Lk+e+++l+++++klk e+
GSVIVT01027040001 285 EREIKRERRKQSNRESARRSRLRKQAETEELALKVESLNTENSVLKSEINRLRENSEKLKLEN 347
89**********************************************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF07777 | 4.3E-33 | 1 | 94 | IPR012900 | G-box binding protein, multifunctional mosaic region |
| Pfam | PF16596 | 4.5E-39 | 132 | 266 | No hit | No description |
| Gene3D | G3DSA:1.20.5.170 | 1.4E-16 | 279 | 344 | No hit | No description |
| SMART | SM00338 | 4.2E-19 | 285 | 349 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF00170 | 5.9E-19 | 285 | 347 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 13.208 | 287 | 350 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 1.13E-10 | 288 | 344 | No hit | No description |
| CDD | cd14702 | 2.08E-20 | 290 | 338 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 292 | 307 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0005829 | Cellular Component | cytosol | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 431 aa Download sequence Send to blast |
MGNEEEGKSP KPEKPTSPPP PDQANIHVYP DWAAMQAYYG PRVTLPPYYN SAMASGHAPH 60 PYIWGPPQPM MPPYGPPYAA IYSPGGVYPH PAVPLGSHSH GHGVQSSPVV SEALAAPPLS 120 IETPAKSSGN TDRGLMKKLK GFDGLAMSIG NGNGESTEGG SDHGLSQSGE TEGSSDGSDG 180 NTAGADQTRR KRSREGTPPI GGDGKTETQA TSAPSAEVNA GSDKVLGVAV PPTSVTGKLA 240 GAVLSPRMST ALELRNPPSV NAKTSPSSIP QPGAMVPSDT WILNEREIKR ERRKQSNRES 300 ARRSRLRKQA ETEELALKVE SLNTENSVLK SEINRLRENS EKLKLENATL MEKLKSAQLE 360 QAEDTHLNKV DDKRVLPVST ENLLSRVNNS GSVDRSTEEE GDMYEKNTNT GAKLHQLLDT 420 SPRADAVAAG * |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 301 | 307 | RRSRLRK |
| 2 | 301 | 308 | RRSRLRKQ |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Vvi.10393 | 0.0 | bud| fruit| inflorescence| leaf | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds to the G-box-like motif (5'-ACGTGGC-3') of the chalcone synthase (CHS) gene promoter. G-box and G-box-like motifs are defined in promoters of certain plant genes which are regulated by such diverse stimuli as light-induction or hormone control. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00318 | DAP | Transfer from AT2G46270 | Download |
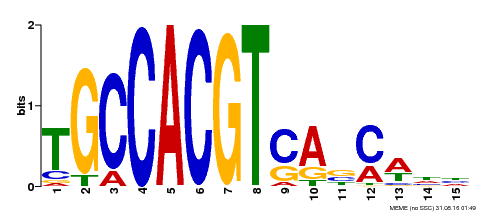
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By light. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AM460965 | 1e-135 | AM460965.1 Vitis vinifera contig VV78X255145.3, whole genome shotgun sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002279966.2 | 0.0 | PREDICTED: common plant regulatory factor 1 isoform X1 | ||||
| Swissprot | Q99089 | 1e-159 | CPRF1_PETCR; Common plant regulatory factor 1 | ||||
| TrEMBL | A0A438JM42 | 0.0 | A0A438JM42_VITVI; Common plant regulatory factor 1 | ||||
| TrEMBL | D7UCK3 | 0.0 | D7UCK3_VITVI; Uncharacterized protein | ||||
| STRING | VIT_15s0046g01440.t01 | 0.0 | (Vitis vinifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP4346 | 11 | 23 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G46270.1 | 2e-92 | G-box binding factor 3 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | GSVIVT01027040001 |
| Entrez Gene | 100248302 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



