 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Eucgr.A00189.1.p | ||||||||
| Common Name | EUGRSUZ_A00189, LOC104440295 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Myrtales; Myrtaceae; Myrtoideae; Eucalypteae; Eucalyptus
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 669aa MW: 72504.4 Da PI: 6.1719 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 91.8 | 6e-29 | 204 | 257 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr++W+ eLH++Fv+av+qL G++kA+Pk+ilelm+v+gLt+e+v+SHLQkYRl
Eucgr.A00189.1.p 204 KPRVVWSVELHQQFVSAVNQL-GIDKAVPKKILELMNVPGLTRENVASHLQKYRL 257
79*******************.********************************8 PP
| |||||||
| 2 | Response_reg | 77.6 | 4.5e-26 | 26 | 134 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTTSEEEEEESTTTHHHHHHHHH CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepklpiivvtahgeeedalealk 91
vl+vdD+p+ +++l+++l+ +y +v++++ ++ al +l++++ +D++l D+ mp+mdG++ll++i e +lp+i+++a + ++ + + +
Eucgr.A00189.1.p 26 VLVVDDDPTCLRILEKMLKTCQY-QVTTCSRSDVALSTLRDNKngFDIVLSDVHMPDMDGFKLLEHIGLEM-DLPVIMMSADDGKQVVMKGVT 116
89*********************.***************888889**********************6654.8******************** PP
TTESEEEESS--HHHHHH CS
Response_reg 92 aGakdflsKpfdpeelvk 109
Ga d+l Kp+ +e+l +
Eucgr.A00189.1.p 117 HGACDYLIKPIRIEALKN 134
**************9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF036392 | 3.8E-176 | 6 | 660 | IPR017053 | Response regulator B-type, plant |
| SuperFamily | SSF52172 | 4.84E-35 | 23 | 147 | IPR011006 | CheY-like superfamily |
| Gene3D | G3DSA:3.40.50.2300 | 8.2E-42 | 23 | 177 | No hit | No description |
| SMART | SM00448 | 3.9E-29 | 24 | 136 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 42.549 | 25 | 140 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 3.1E-23 | 26 | 134 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 5.05E-26 | 27 | 139 | No hit | No description |
| PROSITE profile | PS51294 | 10.732 | 201 | 260 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 4.3E-20 | 201 | 261 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.2E-30 | 203 | 262 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 7.8E-25 | 204 | 257 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 6.1E-8 | 206 | 256 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0071368 | Biological Process | cellular response to cytokinin stimulus | ||||
| GO:0080113 | Biological Process | regulation of seed growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 669 aa Download sequence Send to blast |
MSTLTSSCSW KAGDGVSDQF PAGLRVLVVD DDPTCLRILE KMLKTCQYQV TTCSRSDVAL 60 STLRDNKNGF DIVLSDVHMP DMDGFKLLEH IGLEMDLPVI MMSADDGKQV VMKGVTHGAC 120 DYLIKPIRIE ALKNIWQHVV RKRKNEWKEL EQCGSVEDGD RPQKHLEDVD YSSTANEGSW 180 KSGKRRKEEE DDAEERDDSS TLKKPRVVWS VELHQQFVSA VNQLGIDKAV PKKILELMNV 240 PGLTRENVAS HLQKYRLYLR RLSGVPQHQA GVNSSFIGTQ DAGFGAISSL NGYEIQALAA 300 TGQLAPQSLA TLQAAGLGRS AGKSGISIPM LDQRNLFSFD NPKLRFGEGQ SQQNLGTNSK 360 QANLLHGIPT TMEPKQLVNL QQSAQSFGSM SMPVHTHGAQ SSSLLMPMGQ PQARRHVISE 420 SGSSRPPVLQ ANMGQSIMST GLVGQALGRS GTAENGRGLG HNTILQTSSA MNFPMNQVSD 480 APVSSFPIQS TFPTSKGGYP EDVNSSIIKG PGGGMPSYDI LHDLQQIRSN DWEFQNVGVT 540 FDTSQPGNSV QGNIDTSGLL LVHQGLSSGQ SSNGHNRNIS AAGKSLLSAD GNRQVTTPNV 600 GQPPTSLVDN PVQIKTEGFP DMSYQAVLYP EHYSQEDIMS AFLKQQQGIG PPDSEFEFDG 660 FSVNNIPV* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 2e-22 | 203 | 263 | 4 | 64 | ARR10-B |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-[AG]GATT-3'. Functions as a response regulator involved in His-to-Asp phosphorelay signal transduction system. Phosphorylation of the Asp residue in the receiver domain activates the ability of the protein to promote the transcription of target genes. Could directly activate some type-A response regulators in response to cytokinins. Involved in the expression of nuclear genes for components of mitochondrial complex I. Promotes cytokinin-mediated leaf longevity. Involved in the ethylene signaling pathway in an ETR1-dependent manner and in the cytokinin signaling pathway. {ECO:0000269|PubMed:11370868, ECO:0000269|PubMed:11574878, ECO:0000269|PubMed:15282545, ECO:0000269|PubMed:16407152}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00054 | PBM | Transfer from AT4G16110 | Download |
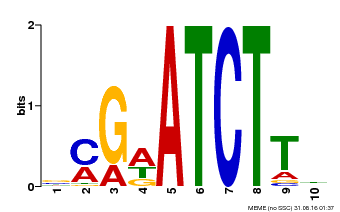
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Eucgr.A00189.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010051575.1 | 0.0 | PREDICTED: two-component response regulator ARR2 | ||||
| Swissprot | Q9ZWJ9 | 0.0 | ARR2_ARATH; Two-component response regulator ARR2 | ||||
| TrEMBL | A0A059DB25 | 0.0 | A0A059DB25_EUCGR; Two-component response regulator | ||||
| STRING | XP_010051575.1 | 0.0 | (Eucalyptus grandis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2403 | 27 | 73 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G16110.1 | 0.0 | response regulator 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Eucgr.A00189.1.p |
| Entrez Gene | 104440295 |



