 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cotton_A_19860_BGI-A2_v1.0 | ||||||||
| Common Name | F383_24767 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 679aa MW: 74406.7 Da PI: 6.5152 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 91.3 | 8.2e-29 | 213 | 266 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr++W+ eLH++Fv av+qL G++kA+Pk+ilelm+v+gLt+e+v+SHLQkYRl
Cotton_A_19860_BGI-A2_v1.0 213 KPRVVWSVELHQQFVAAVNQL-GIDKAVPKKILELMNVPGLTRENVASHLQKYRL 266
79*******************.********************************8 PP
| |||||||
| 2 | Response_reg | 73.3 | 9.8e-25 | 35 | 143 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTTSEEEEEESTT CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepklpiivvtahg 81
vl+vdD+p+ +++l+++l y +v+ + +e al +l+e++ +D++l D+ mp+mdG++ll++i e +lp+i+++a +
Cotton_A_19860_BGI-A2_v1.0 35 VLVVDDDPTCLMILEKMLIACLY-KVTKCNRAETALSKLRENKngYDIVLSDVHMPDMDGFKLLEHIGLEM-DLPVIMMSADD 115
89*****************9999.***************999999**********************6654.8********** PP
THHHHHHHHHTTESEEEESS--HHHHHH CS
Response_reg 82 eeedalealkaGakdflsKpfdpeelvk 109
++ + + + Ga d+l Kp+ +e+l +
Cotton_A_19860_BGI-A2_v1.0 116 GKQVVMKGVTHGACDYLIKPVRIEALKN 143
************************9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF036392 | 8.7E-177 | 9 | 671 | IPR017053 | Response regulator B-type, plant |
| SuperFamily | SSF52172 | 1.85E-34 | 32 | 157 | IPR011006 | CheY-like superfamily |
| Gene3D | G3DSA:3.40.50.2300 | 2.6E-41 | 32 | 174 | No hit | No description |
| SMART | SM00448 | 2.2E-29 | 33 | 145 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 41.725 | 34 | 149 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 4.4E-22 | 35 | 143 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 1.12E-26 | 36 | 148 | No hit | No description |
| PROSITE profile | PS51294 | 11.048 | 210 | 269 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 3.58E-20 | 210 | 270 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.5E-30 | 212 | 271 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 9.7E-25 | 213 | 266 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 9.8E-8 | 215 | 265 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0071368 | Biological Process | cellular response to cytokinin stimulus | ||||
| GO:0080113 | Biological Process | regulation of seed growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 679 aa Download sequence Send to blast |
MNVSSVKGSM SISSSTSTWK AGDTISDQFP AGLRVLVVDD DPTCLMILEK MLIACLYKVT 60 KCNRAETALS KLRENKNGYD IVLSDVHMPD MDGFKLLEHI GLEMDLPVIM MSADDGKQVV 120 MKGVTHGACD YLIKPVRIEA LKNIWQHVVR KRKNEWKEFE QSGSVEEGDR QPKQSDDADY 180 SSSANEGNWK GSKKRKDEEE ETDERDDTST LKKPRVVWSV ELHQQFVAAV NQLGIDKAVP 240 KKILELMNVP GLTRENVASH LQKYRLYLRR LSGVSQHQSN LSTIISAQDP TFGSLSSLSG 300 LDLQTLAATG QLPAQSLARL QAAGLGRATA KSGIPITLVD QRNIFSFENP KLRFGEGQQQ 360 HMTNKQQVNL LHGIPTTMEP KQLVSLRHAA QSIGNMNMQV PPHGAQNSQN NPLLMQMGQQ 420 QQQSRGQILV DSTINHAPRL SSSMGQPILS NGIATNVSSR NGIPENIRAP GYSQTPSMLN 480 FPVNHASELP GNSFALGSTP GVSILTSKGA FQEDVNSEIK GSGGLMPSYD VFNDLNQHKP 540 QSWELQNVGM AFDTSQHSNS LQGNLDLTQS ALGQQGFSSG QMNGHNRSAA VANKAMFSTG 600 DVKELGSAQN VNQHLNNLLV DNTIRIKSER VCDTSPANIF PDHFGQDDLM SALLKQQESV 660 ASSENEFDFD GYSMNNIPV |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 7e-22 | 212 | 272 | 4 | 64 | ARR10-B |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-[AG]GATT-3'. Functions as a response regulator involved in His-to-Asp phosphorelay signal transduction system. Phosphorylation of the Asp residue in the receiver domain activates the ability of the protein to promote the transcription of target genes. Could directly activate some type-A response regulators in response to cytokinins. Involved in the expression of nuclear genes for components of mitochondrial complex I. Promotes cytokinin-mediated leaf longevity. Involved in the ethylene signaling pathway in an ETR1-dependent manner and in the cytokinin signaling pathway. {ECO:0000269|PubMed:11370868, ECO:0000269|PubMed:11574878, ECO:0000269|PubMed:15282545, ECO:0000269|PubMed:16407152}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00054 | PBM | Transfer from AT4G16110 | Download |
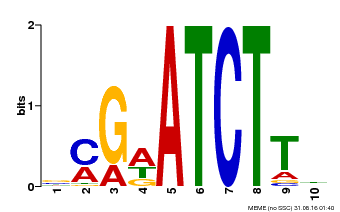
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | GQ393465 | 6e-83 | GQ393465.1 Gossypium hirsutum cultivar Deltapine 33 B clone MONCS0430 SSR marker CGR5416 genomic sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017649032.1 | 0.0 | PREDICTED: two-component response regulator ARR2-like isoform X1 | ||||
| Swissprot | Q9ZWJ9 | 0.0 | ARR2_ARATH; Two-component response regulator ARR2 | ||||
| TrEMBL | A0A0B0MT30 | 0.0 | A0A0B0MT30_GOSAR; Two-component response regulator | ||||
| STRING | Gorai.011G196400.1 | 0.0 | (Gossypium raimondii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2403 | 27 | 73 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G16110.1 | 0.0 | response regulator 2 | ||||



