 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Ciclev10000183m | ||||||||
| Common Name | CICLE_v10000183mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Rutaceae; Aurantioideae; Citrus
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 947aa MW: 104767 Da PI: 5.6923 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 77.3 | 1.7e-24 | 147 | 248 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SS CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgr 87
f+k+lt sd++++g +++p++ ae+ ++++ + ++l+++d++ ++W++++iyr++++r++lt+GW+ Fv +++L++gD+v+F ++
Ciclev10000183m 147 FCKTLTASDTSTHGGFSVPRRAAEKLfppldytMQPP-T-QELVVRDLHDNTWTFRHIYRGQPKRHLLTTGWSLFVGSKRLRAGDSVLFI--RD 236
99*********************999*****954444.4.38************************************************..45 PP
SEE..EEEEE-S CS
B3 88 sefelvvkvfrk 99
++ +l+v+v+r+
Ciclev10000183m 237 EKSQLMVGVRRA 248
77778*****97 PP
| |||||||
| 2 | Auxin_resp | 115.5 | 4.8e-38 | 273 | 356 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalk.vkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
aahaas++s F+++YnPra++s+Fv++++k++k+++ +++svGmRf m+fete+s +rr++Gt+vg+sdldp+rWp+SkWr+L+
Ciclev10000183m 273 AAHAASNRSQFTIFYNPRACPSDFVIPLAKYRKSVYgTQMSVGMRFGMMFETEESGKRRYMGTIVGISDLDPLRWPGSKWRNLQ 356
79**********************************9*********************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 7.72E-47 | 134 | 276 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 1.0E-41 | 140 | 261 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 4.29E-23 | 146 | 247 | No hit | No description |
| Pfam | PF02362 | 5.4E-22 | 147 | 248 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 3.0E-25 | 147 | 249 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 13.211 | 147 | 249 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 4.3E-33 | 273 | 356 | IPR010525 | Auxin response factor |
| SuperFamily | SSF54277 | 5.4E-10 | 831 | 911 | No hit | No description |
| PROSITE profile | PS51745 | 25.619 | 832 | 916 | IPR000270 | PB1 domain |
| Pfam | PF02309 | 3.2E-10 | 832 | 917 | IPR033389 | AUX/IAA domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0009908 | Biological Process | flower development | ||||
| GO:0009942 | Biological Process | longitudinal axis specification | ||||
| GO:0010305 | Biological Process | leaf vascular tissue pattern formation | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:0048507 | Biological Process | meristem development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016020 | Cellular Component | membrane | ||||
| GO:0042802 | Molecular Function | identical protein binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 947 aa Download sequence Send to blast |
MGSVEEKIKA GGLVIRAQTT LLEEMKLLKE MQDQSGARKA INSELWHACA GPLVFLPQVG 60 SLVYYFPQGH SEQVAASTKR SATSQIPNYP NLPSQLLCQV HNVTLHADKD TDEIYAQMSL 120 QPVNSEKDVF PIPDFGLKPS KHPSEFFCKT LTASDTSTHG GFSVPRRAAE KLFPPLDYTM 180 QPPTQELVVR DLHDNTWTFR HIYRGQPKRH LLTTGWSLFV GSKRLRAGDS VLFIRDEKSQ 240 LMVGVRRANR QQTALPSSVL SADSMHIGVL AAAAHAASNR SQFTIFYNPR ACPSDFVIPL 300 AKYRKSVYGT QMSVGMRFGM MFETEESGKR RYMGTIVGIS DLDPLRWPGS KWRNLQVEWD 360 EPGCSDKQKR VSPWEIETPE SLFIFPSLTS GLKRPFHSGI LATETEWGSL IKRPLACPEI 420 APGVMPYSSI SNLCSEQLIK MMLKPQLVNN PGSFAASSLQ ETSGAKGAHL EEVKTLQSTI 480 NQKPRLVPSE MNRIDNQNCS QICLNQADTV NSSLSRIHIP EKPHPPSKCE KQAPPGMNTD 540 HLKSEPRQSI EQSSNLTSAA DCSMEKPSGP LNPQNLVNQH AFHNQNEGLL QLQSSWPMQS 600 QLESVFQAQQ INVPQSDSTA HSGSLPILDT DEWMSHTSCN SLAGTYNRSP GPLPMFGLQE 660 PSTMLPEVIN PSLSFPGQEM WDHQLNNLRF LSPVDPLTSF TQQDHCSLNS SGLRDLSDES 720 NNQSGIYSCL NIDVSNGGST MIDHSVSSAI LDEFCTLKDA NFQNPPDCLM NTFSSSQDVQ 780 SQITSASLAD SQAFSRQDFP DNSGGTSSSN VDFDESSLLQ NTSWQPVVPP MRTYTKVQKT 840 GSVGRSIDVT NFKNYDELCS AIERMFGLEG LLNDPRGTEW KLVYVDYEND VLLVGDDPWE 900 EFVGCVRCIR ILSPQEVQQM SEEGMKLLNS AAMQGIDCTK PEGGRA* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldu_A | 0.0 | 1 | 381 | 5 | 392 | Auxin response factor 5 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Ccl.22384 | 6e-46 | abscission zone | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: In early embryo and during organ development. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in the whole plant with a lower expression in leaves. Detected in embryo axis, provascular tissues, procambium and some differentiated vascular regions of mature organs. {ECO:0000269|PubMed:10476078}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). Seems to act as transcriptional activator. Formation of heterodimers with Aux/IAA proteins may alter their ability to modulate early auxin response genes expression. Mediates embryo axis formation and vascular tissues differentiation. Functionally redundant with ARF7. May be necessary to counteract AMP1 activity. {ECO:0000269|PubMed:12036261, ECO:0000269|PubMed:14973283, ECO:0000269|PubMed:17553903}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00153 | DAP | Transfer from AT1G19850 | Download |
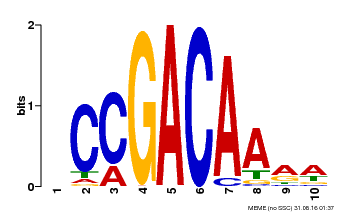
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024039097.1 | 0.0 | auxin response factor 5 isoform X1 | ||||
| Swissprot | P93024 | 0.0 | ARFE_ARATH; Auxin response factor 5 | ||||
| TrEMBL | V4TFD4 | 0.0 | V4TFD4_9ROSI; Auxin response factor | ||||
| STRING | XP_006435146.1 | 0.0 | (Citrus clementina) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM5808 | 27 | 46 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G19850.1 | 0.0 | ARF family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Ciclev10000183m |
| Entrez Gene | 18042398 |



