 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Bv3_048080_hfrw.t2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Chenopodiaceae; Betoideae; Beta
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 649aa MW: 71274 Da PI: 6.1415 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 91.4 | 7.8e-29 | 191 | 244 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr++W+ eLH++Fv av+qL G++kA+Pk+ilelm+v+gLt+e+v+SHLQkYRl
Bv3_048080_hfrw.t2 191 KPRVVWSVELHQQFVAAVNQL-GIDKAVPKKILELMNVPGLTRENVASHLQKYRL 244
79*******************.********************************8 PP
| |||||||
| 2 | Response_reg | 75.2 | 2.5e-25 | 15 | 123 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTTSEEEEEESTTTHHHHHHH CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepklpiivvtahgeeedalea 89
vl+vdD+p+ + +l+++l+ +y ev+ + +e al +l+e++ +D+++ D+ mp+mdG++ll++ e +lp+i+++a +++ + +
Bv3_048080_hfrw.t2 15 VLVVDDDPTCLTILEKMLRTCRY-EVTKTNRAEVALSWLRENRngFDIVISDVHMPDMDGFKLLEHVGLEM-DLPVIMMSADDSKGVVMKG 103
89*********************.***************99999***********************6644.8****************** PP
HHTTESEEEESS--HHHHHH CS
Response_reg 90 lkaGakdflsKpfdpeelvk 109
+ Ga d+l Kp+ +e+l +
Bv3_048080_hfrw.t2 104 VTHGACDYLIKPVRIEALKN 123
****************9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF036392 | 2.2E-174 | 1 | 638 | IPR017053 | Response regulator B-type, plant |
| Gene3D | G3DSA:3.40.50.2300 | 2.2E-41 | 12 | 152 | No hit | No description |
| SuperFamily | SSF52172 | 2.13E-34 | 12 | 135 | IPR011006 | CheY-like superfamily |
| SMART | SM00448 | 1.2E-28 | 13 | 125 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 41.592 | 14 | 129 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 1.5E-22 | 15 | 123 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 3.66E-25 | 16 | 128 | No hit | No description |
| PROSITE profile | PS51294 | 11.048 | 188 | 247 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 3.05E-20 | 188 | 248 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.1E-30 | 190 | 249 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 9.2E-25 | 191 | 244 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 9.2E-8 | 193 | 243 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0071368 | Biological Process | cellular response to cytokinin stimulus | ||||
| GO:0080113 | Biological Process | regulation of seed growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 649 aa Download sequence Send to blast |
MADSSGDQFP VGLRVLVVDD DPTCLTILEK MLRTCRYEVT KTNRAEVALS WLRENRNGFD 60 IVISDVHMPD MDGFKLLEHV GLEMDLPVIM MSADDSKGVV MKGVTHGACD YLIKPVRIEA 120 LKNIWQHVVR KKKNEYRDVE QSGSWDEGDR QLKQDDAVSS PANEGNWKNA KRKSGEDDES 180 EDKDDTTTLK KPRVVWSVEL HQQFVAAVNQ LGIDKAVPKK ILELMNVPGL TRENVASHLQ 240 KYRLYLRRLS GVSQHQGGLN NSFMPQDPSF STITQLGGID LQTLAATGQL SAQTLAAYTR 300 LPPTIKPGMS MPFVDQRNLF SFENSKLRYG EGQQSQISNA NKQMNLLHGI PTTMEPKQLA 360 GMNQSAQTLG SMNMQATASS SHQNSSLLMH PMVPQQSRGH ISNESVSSQV SRIQPSIGQP 420 LHSNGTASAV LSRNSMGYDP VSQAGPVVDF SVNQISELPG NGFPLGSTPG MTSISSKGFN 480 QEEVSSDIKV TRGFVGSYDL FSELQHKSQD WQLQNPNMSY AGSSQHVTSI QNGVNVAPSI 540 MVNQSYVPAQ KNEQNGHAMS GKLMYSTGLE NQHVGMQNMN HNLNSVLAHS SSRVKAESVS 600 DVVNPGANLF SEPYGQEDLM SALLKQQEGI GSGEFDFDGY TLDNIPVNM |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 2e-22 | 187 | 250 | 1 | 64 | ARR10-B |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-[AG]GATT-3'. Functions as a response regulator involved in His-to-Asp phosphorelay signal transduction system. Phosphorylation of the Asp residue in the receiver domain activates the ability of the protein to promote the transcription of target genes. Could directly activate some type-A response regulators in response to cytokinins. Involved in the expression of nuclear genes for components of mitochondrial complex I. Promotes cytokinin-mediated leaf longevity. Involved in the ethylene signaling pathway in an ETR1-dependent manner and in the cytokinin signaling pathway. {ECO:0000269|PubMed:11370868, ECO:0000269|PubMed:11574878, ECO:0000269|PubMed:15282545, ECO:0000269|PubMed:16407152}. | |||||
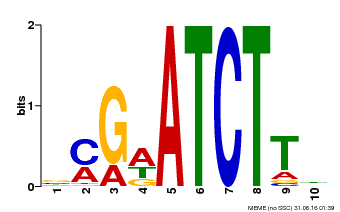
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00054 | PBM | Transfer from AT4G16110 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010670915.1 | 0.0 | PREDICTED: two-component response regulator ARR2 isoform X2 | ||||
| Swissprot | Q9ZWJ9 | 0.0 | ARR2_ARATH; Two-component response regulator ARR2 | ||||
| TrEMBL | A0A0K9RPY9 | 0.0 | A0A0K9RPY9_SPIOL; Two-component response regulator | ||||
| STRING | XP_010670914.1 | 0.0 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G16110.1 | 0.0 | response regulator 2 | ||||



