 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Aradu.H2IIR | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Dalbergieae; Arachis
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 1007aa MW: 110877 Da PI: 6.3783 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 131.6 | 2.7e-41 | 149 | 226 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+C adls+ak+yhrrhkvCe+hska+++lv + +qrfCqqCsrfh+l+efDe+krsCrrrLa+hn+rrrk+++
Aradu.H2IIR 149 VCQVEDCGADLSSAKDYHRRHKVCEMHSKATKALVGNAMQRFCQQCSRFHMLQEFDEGKRSCRRRLAGHNKRRRKTNN 226
6**************************************************************************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 1.1E-33 | 143 | 211 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.129 | 147 | 224 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 3.79E-38 | 148 | 228 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 5.9E-30 | 150 | 223 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF48403 | 7.46E-7 | 779 | 891 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 7.79E-7 | 781 | 892 | No hit | No description |
| Gene3D | G3DSA:1.25.40.20 | 9.3E-7 | 781 | 895 | IPR020683 | Ankyrin repeat-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1007 aa Download sequence Send to blast |
MEARFGTEAL QFYAMGGSSV GKRSMEWDLN DWKWDGDLFI ARPLNPGPEH RQFFPAGTRI 60 PVAGGPSNSN SSSSCSDEVD LGIRGGNKEG DKKRRVVVLE DDALNEESGT LSLKLGGGVG 120 GHGGAQIGSW EGVNGKKSRV AGGNSNRAVC QVEDCGADLS SAKDYHRRHK VCEMHSKATK 180 ALVGNAMQRF CQQCSRFHML QEFDEGKRSC RRRLAGHNKR RRKTNNDAVP NATSLNDDQT 240 SSYLLISLLK ILSNMHSDRS DQTTDQDLLT HLLRSLASQN GDQGGKNLSN LLAQPENLLK 300 EGSSSGKSEM VSTLFSNGSQ GSPSVIQQHQ AISVAELQHQ VMHAHDIMPT DQQIMSSTKP 360 SASNSPPTYS EARDSTAGQT KINNFDLNDI YIDSDDGIED IEKLPVSANL GTSSLEYPWT 420 QHDSHQSSPP QTSGNSDSAS AQSPSSSSGE AQSRTDRIVF KLFGKEPNDF PLVLRAQILD 480 WLSHSPTDME SYIRPGCIVL TIYLRQSEAM WEELCYDLTS SLSRLLDVSD VDFWRTGWVH 540 IRVQHQLAFV FNGQVVIDTS LPFRSNNYGK ILSVSPIAVP ASKTAHFSVK GINLNRPATR 600 LLCALEGNYL SCEDAHESMD QGSKELEELQ CIQFSCSVPV INGRGFIEIE DQGLSSSYFP 660 FIVAEEDVCS EICVLEPLIE VSDIDPDNEG TGKIKAKNQA MDFIHEMGWL LHRSQLRSRM 720 VHLNSSVELF PLKRFKWLME FSVDRDWCAV VKKLLNLLLS GTVGTGDHQS LHLALSEMGL 780 LHKAVRRNSR QLVELLLRYV PENISDKLGH EDMALVGGEN QSFLFRPDAA GPAGLTPLHI 840 AAGKDGSEEV LDALTNDPCM VGIEAWKSAR DSTGSTPEDY ARLRGHYAYI HLVQKKINKR 900 QGGAHVVVDI PSNLTGFNTN QKQNETTSFD IVGKAEGRSA QKQCKLCDNK LSCRAVVGKS 960 LAYRPAMLSM VAIAAVCVCV ALLFKSSPEV LYVFRPFRWE SLEYGTS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 1e-30 | 141 | 223 | 2 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Binds specifically to the 5'-GTAC-3' core sequence. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:16095614}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
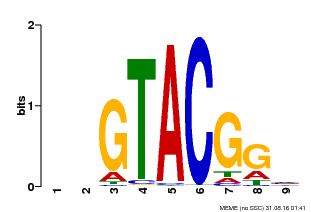
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Aradu.H2IIR |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KF312231 | 0.0 | KF312231.1 Arachis hypogaea squamosa promoter-binding-like protein (SPL12) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015968830.1 | 0.0 | squamosa promoter-binding-like protein 1 | ||||
| Refseq | XP_015968831.1 | 0.0 | squamosa promoter-binding-like protein 1 | ||||
| Swissprot | Q9SMX9 | 0.0 | SPL1_ARATH; Squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | A0A445CMW3 | 0.0 | A0A445CMW3_ARAHY; Uncharacterized protein | ||||
| STRING | GLYMA02G01160.1 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1539 | 34 | 95 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



