 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 903325 | ||||||||
| Common Name | ARALYDRAFT_903325 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 351aa MW: 38921.8 Da PI: 4.3638 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 55.1 | 1.9e-17 | 81 | 131 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkr.krfslgkfgtaeeAakaaiaarkkleg 55
++y+GVr++ +g+WvAeIr+p g r +r +lg+f + +eAa a+++a+k+++g
903325 81 CDYRGVRQRT-WGKWVAEIREP---G-RgARLWLGTFSSSYEAALAYDEAAKAIYG 131
68*****999.**********8...4.45***********************9987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 7.27E-31 | 81 | 140 | No hit | No description |
| Pfam | PF00847 | 4.9E-13 | 81 | 131 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 4.6E-36 | 82 | 145 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 20.916 | 82 | 139 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 5.75E-19 | 83 | 139 | IPR016177 | DNA-binding domain |
| PRINTS | PR00367 | 7.8E-9 | 83 | 94 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 2.2E-28 | 83 | 140 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 7.8E-9 | 105 | 121 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0010286 | Biological Process | heat acclimation | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 351 aa Download sequence Send to blast |
MGVLEHVANL ASMPFDSPRK RKSRGTRDVA EILRQWREYN EQTEADSCID GGVPKPVRNP 60 PPKGSRKGCM KGKGGPENGI CDYRGVRQRT WGKWVAEIRE PGRGARLWLG TFSSSYEAAL 120 AYDEAAKAIY GQSARLNLPE ITNRSSSTAA TVSGSVTAFS DESEVCARED TNERSGFGQV 180 KLEDCNDEYV LLDSSQCIKE ELKVKEEVME EHNSAVGFGI GQDPKREILD AWLMGNGNEQ 240 EPLEFGVDET FDINELLGIL DDNNVSGQET MQNQVDRQPN FSYQTQFQDA NLLGSLNPME 300 IAHPGVDYGY PYVQPSEMEN NGIDLDHHRF NDLDIQDLDF GGEKDVHGST * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1gcc_A | 4e-17 | 83 | 139 | 3 | 60 | ETHYLENE RESPONSIVE ELEMENT BINDING FACTOR 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-[AG]CCGAC-3'. Binding to the C-repeat/DRE element mediates high salinity- and abscisic acid-inducible transcription. {ECO:0000269|PubMed:11798174}. | |||||
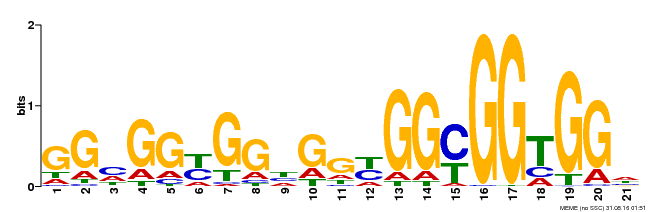
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00302 | DAP | Transfer from AT2G40340 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 903325 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By high-salt stress and abscisic acid (ABA) treatment. {ECO:0000269|PubMed:11798174}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC007020 | 0.0 | AC007020.5 Arabidopsis thaliana chromosome 2 clone T3G21 map g4514, complete sequence. | |||
| GenBank | AF085279 | 0.0 | AF085279.1 Arabidopsis thaliana ABA-regulated gene cluster, complete sequence. | |||
| GenBank | CP002685 | 0.0 | CP002685.1 Arabidopsis thaliana chromosome 2, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002881716.1 | 0.0 | dehydration-responsive element-binding protein 2C isoform X1 | ||||
| Swissprot | Q8LFR2 | 0.0 | DRE2C_ARATH; Dehydration-responsive element-binding protein 2C | ||||
| TrEMBL | D7LF31 | 0.0 | D7LF31_ARALL; Uncharacterized protein | ||||
| STRING | scaffold_402826.1 | 0.0 | (Arabidopsis lyrata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM8532 | 25 | 38 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G40340.1 | 0.0 | ERF family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 903325 |
| Entrez Gene | 9317782 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



