 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 676781038 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Sisymbrieae; Sisymbrium
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 453aa MW: 50038.3 Da PI: 9.7895 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 44.7 | 2.8e-14 | 375 | 426 | 5 | 55 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH.HHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkke.leelkke 55
+r++r++kNRe+A rsR+RK+a++ eLe +v++L++ N++L+ + +e ++k+
676781038 375 RRQKRMIKNRESAARSRARKQAYTLELEAEVAQLKELNEELQRKqVEIMEKQ 426
69*************************************9975404434444 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 1.2E-13 | 371 | 436 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 10.507 | 373 | 418 | IPR004827 | Basic-leucine zipper domain |
| Gene3D | G3DSA:1.20.5.170 | 1.9E-13 | 375 | 430 | No hit | No description |
| CDD | cd14707 | 1.16E-26 | 375 | 429 | No hit | No description |
| SuperFamily | SSF57959 | 9.14E-10 | 375 | 426 | No hit | No description |
| Pfam | PF00170 | 2.7E-12 | 375 | 431 | IPR004827 | Basic-leucine zipper domain |
| PROSITE pattern | PS00036 | 0 | 378 | 393 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009738 | Biological Process | abscisic acid-activated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0000976 | Molecular Function | transcription regulatory region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 453 aa Download sequence Send to blast |
MGSRMNLKNF VDGVSDEVIY TTKQPALGTA ALPLTRQNSV FSLTFDEFQN SWGGGIGKDF 60 GSMNMDELLK NIWTAEENHS MMTNNTTFNN VNNGSSGNAV TNVGNNNGGL SVGVGGGEVG 120 GFFTGGGLQR QRSLTLPRTI SQKRVDDVWK ELMKEDDTGN GVANHGASGI PQRQQTLGEM 180 TLEEFLVRAG VVREEPQPVE RLDNFNGGFY GFGTNAGLGS ARNNGFGPNQ PHDLSGNGAV 240 RPDLLTTQTQ PLQVQQPQKV HQQQQLIQKP ETPFAKQTTI TFTNTVDANN RPQPATQYQE 300 VKPSILGVRD LPLNNNLLQA VDYKTGVTVA AVSPGSQMSP DLTPKSTMDT TFSPVPYMFG 360 RVRKTGAVLE KVIERRQKRM IKNRESAARS RARKQAYTLE LEAEVAQLKE LNEELQRKQV 420 EIMEKQKKQL LEPMRQPWGC KRQCLRRTLT GPW |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds to the ABA-responsive element (ABRE). Mediates stress-responsive ABA signaling. {ECO:0000269|PubMed:11884679, ECO:0000269|PubMed:15361142}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00038 | PBM | Transfer from AT4G34000 | Download |
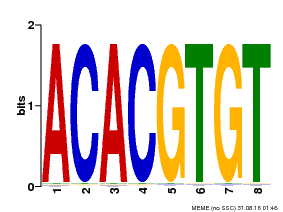
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 676781038 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by drought, salt, abscisic acid (ABA). {ECO:0000269|PubMed:10636868, ECO:0000269|PubMed:16284313}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK353408 | 0.0 | AK353408.1 Thellungiella halophila mRNA, complete cds, clone: RTFL01-30-J01. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006412286.1 | 0.0 | ABSCISIC ACID-INSENSITIVE 5-like protein 6 | ||||
| Refseq | XP_024005691.1 | 0.0 | ABSCISIC ACID-INSENSITIVE 5-like protein 6 | ||||
| Refseq | XP_024005692.1 | 0.0 | ABSCISIC ACID-INSENSITIVE 5-like protein 6 | ||||
| Swissprot | Q9M7Q3 | 0.0 | AI5L6_ARATH; ABSCISIC ACID-INSENSITIVE 5-like protein 6 | ||||
| TrEMBL | E4MY11 | 0.0 | E4MY11_EUTHA; mRNA, clone: RTFL01-30-J01 | ||||
| TrEMBL | V4MBG2 | 0.0 | V4MBG2_EUTSA; Uncharacterized protein | ||||
| STRING | XP_006412286.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM734 | 27 | 129 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G34000.2 | 0.0 | abscisic acid responsive elements-binding factor 3 | ||||



