 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 491272 | ||||||||
| Common Name | ARALYDRAFT_491272 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 448aa MW: 48836.9 Da PI: 8.4759 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 43.2 | 8.9e-14 | 367 | 417 | 5 | 54 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH.HHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkke.leelkk 54
+r++r++kNRe+A rsR+RK+a++ eLe ++++L++ N++L+ + +e ++k
491272 367 RRQKRMIKNRESAARSRARKQAYTMELEAEIAQLKELNEELQRKqVEIMEK 417
69*************************************996540333344 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 1.1E-13 | 363 | 428 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 10.45 | 365 | 410 | IPR004827 | Basic-leucine zipper domain |
| Gene3D | G3DSA:1.20.5.170 | 3.5E-13 | 367 | 423 | No hit | No description |
| CDD | cd14707 | 1.34E-27 | 367 | 421 | No hit | No description |
| Pfam | PF00170 | 1.6E-11 | 367 | 424 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 9.38E-10 | 367 | 420 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 370 | 385 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009738 | Biological Process | abscisic acid-activated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0000976 | Molecular Function | transcription regulatory region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 448 aa Download sequence Send to blast |
MGSRLNFKSF VDGVSEQPTV GTSLPLTRQN SVFSLTFDEF QNSWGGGIGK DFGSMNMDEL 60 LKNIWTAEES HSMMGNNTSF NNINNGNSGN TVINGGGNNN GGLAVGVGGE SGGFFTGGSL 120 QRQGSITLPR TISQKRVDDV WKELMEEDDT GNGVGNGGTS GIPQRQQTLG EMTLEEFLVR 180 AGVVREEPQP VESVTNFHGG FYGFGSNGGL GTAINGFGAN QPHDLSGNGA VMRPDLLTAQ 240 TQPLQMQQPQ TVQQPQQLIQ KQERPFPKQT TIAFSNTVDA VNRSQPATQC QEVKPSILGI 300 HNHPMNNNLL QAVEFKTGVT VAAVSPGSQM SPDLTPKSAL DASLSPVPYM FGRVRKTGAV 360 LEKVIERRQK RMIKNRESAA RSRARKQAYT MELEAEIAQL KELNEELQRK QVEIMEKQKN 420 QLLEPLRQPW GMGCKRQCLR RTLTGPW* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds to the ABA-responsive element (ABRE). Mediates stress-responsive ABA signaling. {ECO:0000269|PubMed:11884679, ECO:0000269|PubMed:15361142}. | |||||
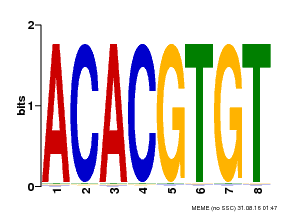
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00038 | PBM | Transfer from AT4G34000 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 491272 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by drought, salt, abscisic acid (ABA). {ECO:0000269|PubMed:10636868, ECO:0000269|PubMed:16284313}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF093546 | 0.0 | AF093546.1 Arabidopsis thaliana clone 11 abscisic acid responsive elements-binding factor (ABRE) mRNA, complete cds. | |||
| GenBank | AK175851 | 0.0 | AK175851.1 Arabidopsis thaliana mRNA for abscisic acid responsive elements-binding factor (ABRE/ABF3), complete cds, clone: RAFL22-47-B14. | |||
| GenBank | AY054605 | 0.0 | AY054605.1 Arabidopsis thaliana Unknown protein (F17I5.190:F17I5.200) mRNA, complete cds. | |||
| GenBank | AY081467 | 0.0 | AY081467.1 Arabidopsis thaliana unknown protein (F17I5.190:F17I5.200) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020874468.1 | 0.0 | ABSCISIC ACID-INSENSITIVE 5-like protein 6 | ||||
| Refseq | XP_020874469.1 | 0.0 | ABSCISIC ACID-INSENSITIVE 5-like protein 6 | ||||
| Refseq | XP_020874470.1 | 0.0 | ABSCISIC ACID-INSENSITIVE 5-like protein 6 | ||||
| Refseq | XP_020874471.1 | 0.0 | ABSCISIC ACID-INSENSITIVE 5-like protein 6 | ||||
| Swissprot | Q9M7Q3 | 0.0 | AI5L6_ARATH; ABSCISIC ACID-INSENSITIVE 5-like protein 6 | ||||
| TrEMBL | D7MFS7 | 0.0 | D7MFS7_ARALL; Uncharacterized protein | ||||
| STRING | fgenesh2_kg.7__705__AT4G34000.1 | 0.0 | (Arabidopsis lyrata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM734 | 27 | 129 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G34000.2 | 0.0 | abscisic acid responsive elements-binding factor 3 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 491272 |
| Entrez Gene | 9303211 |



