 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zpz_sc03807.1.g00060.1.am.mk | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 407aa MW: 43761.9 Da PI: 11.0921 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 41.8 | 2e-13 | 164 | 210 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksLq 55
+h e+Er+RR+++N++f Lr ++Pk + K++Ka+ L A+ YI++L+
Zpz_sc03807.1.g00060.1.am.mk 164 NHVEAERQRREKLNQRFYALRAVVPKiS------KMDKASLLSDAIAYIQELE 210
799***********************66......****************996 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 4.8E-9 | 2 | 64 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 17.047 | 160 | 209 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 2.5E-16 | 163 | 213 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 4.63E-14 | 163 | 214 | No hit | No description |
| SuperFamily | SSF47459 | 1.7E-16 | 163 | 215 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 6.0E-11 | 164 | 210 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 8.1E-16 | 166 | 215 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 407 aa Download sequence Send to blast |
LASRRHAWAA VDPCGPAPGW FVRASLAQAA GLRTVVFLPC NGGVLELGSV VPVRETPEAL 60 RAIEAALAIT PSAPEESMRI FGKDLSRRGQ LPAAAQANWA PPQLGGQSSP DDRKKDAVEQ 120 KPRDPPPKRV DFTKLEPNGQ AVVEERKPRK RGRKPANGRE EPLNHVEAER QRREKLNQRF 180 YALRAVVPKI SKMDKASLLS DAIAYIQELE DRLRGGAKSP PAVEVKAMQD DVVLRVTTPL 240 HAHPASRVFH AIRDARLSVV ASELAVADDA VTHTLMLRSP GPERLTAEAV LAAMSRATEG 300 RDTPSPPASV AYAPFTLTTG RSCAFSAPSK SFLLRGRHPC DVWEPRRPRV VVKWKSGGGA 360 RASDGSRGAS ACYARGRGKG RRGDGTRLGL GGAGVHGGSA ERRRPIG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 2e-24 | 158 | 213 | 2 | 57 | Transcription factor MYC2 |
| 5gnj_B | 2e-24 | 158 | 213 | 2 | 57 | Transcription factor MYC2 |
| 5gnj_E | 2e-24 | 158 | 213 | 2 | 57 | Transcription factor MYC2 |
| 5gnj_F | 2e-24 | 158 | 213 | 2 | 57 | Transcription factor MYC2 |
| 5gnj_G | 2e-24 | 158 | 213 | 2 | 57 | Transcription factor MYC2 |
| 5gnj_I | 2e-24 | 158 | 213 | 2 | 57 | Transcription factor MYC2 |
| 5gnj_M | 2e-24 | 158 | 213 | 2 | 57 | Transcription factor MYC2 |
| 5gnj_N | 2e-24 | 158 | 213 | 2 | 57 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 145 | 153 | RKPRKRGRK |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00099 | PBM | Transfer from AT4G16430 | Download |
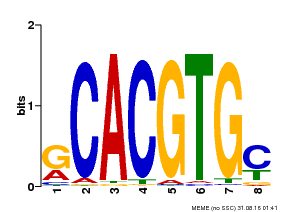
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Zpz_sc03807.1.g00060.1.am.mk |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK064946 | 1e-104 | AK064946.1 Oryza sativa Japonica Group cDNA clone:J013000P16, full insert sequence. | |||
| GenBank | AP003023 | 1e-104 | AP003023.2 Oryza sativa Japonica Group genomic DNA, chromosome 1, PAC clone:P0684B02. | |||
| GenBank | AP003381 | 1e-104 | AP003381.3 Oryza sativa Japonica Group genomic DNA, chromosome 1, PAC clone:P0692C11. | |||
| GenBank | AP014957 | 1e-104 | AP014957.1 Oryza sativa Japonica Group DNA, chromosome 1, cultivar: Nipponbare, complete sequence. | |||
| GenBank | CP012609 | 1e-104 | CP012609.1 Oryza sativa Indica Group cultivar RP Bio-226 chromosome 1 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002456225.1 | 1e-128 | transcription factor bHLH13 | ||||
| TrEMBL | C5XHM5 | 1e-126 | C5XHM5_SORBI; Uncharacterized protein | ||||
| STRING | Sb03g032420.1 | 1e-127 | (Sorghum bicolor) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP7385 | 29 | 44 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G46510.1 | 7e-44 | ABA-inducible BHLH-type transcription factor | ||||



