 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zpz_sc01411.1.g00210.1.am.mk | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 493aa MW: 54134.5 Da PI: 11.5488 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 56.6 | 6.6e-18 | 321 | 371 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
+ ykGVr++ grWv+e+r+p g ++r++lg+f tae+Aa+a++ a+++l+g
Zpz_sc01411.1.g00210.1.am.mk 321 PVYKGVRRRN-PGRWVCEVREP--HG-KQRIWLGTFETAEMAARAHDVAALALRG 371
68*****887.9******9996..33.5*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF00847 | 2.7E-13 | 321 | 371 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 1.83E-20 | 322 | 380 | IPR016177 | DNA-binding domain |
| PROSITE profile | PS51032 | 21.694 | 322 | 379 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 1.4E-26 | 322 | 385 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 3.6E-29 | 322 | 381 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 9.8E-8 | 323 | 334 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 4.42E-25 | 323 | 381 | No hit | No description |
| PRINTS | PR00367 | 9.8E-8 | 345 | 361 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009631 | Biological Process | cold acclimation | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 493 aa Download sequence Send to blast |
MMHAMPKTLV GGGKLGKAES ARDGAEDRDG VRGEAATRAR VADEERTVHG GGRRREQKGG 60 RRDGDRSRRH LRRKPPNQRS RLSVRSKGKP LETGARAPQD TASRRPVRRH WRSISNPRDR 120 VQRNYRSTVT LEGSSARPWT HRLRWSGVYD AQRIRFRQSP PSRKRERERE RERARAPGWG 180 EAQAGGRRPM TPRCARPNRS SAPRAPAGSD PTNLRSPLAR EPVPRVLDPS SRANLAGISG 240 AATRRSSAHG HVALDAPSTL PSRRSIRSIA MDVGALSSDY SSGTPSPVGG SDGSSASYMT 300 VSSAPPKRRA GRTKFKETRH PVYKGVRRRN PGRWVCEVRE PHGKQRIWLG TFETAEMAAR 360 AHDVAALALR GLGACLNFAD SPRLLRVPPL GAGHDEIRRA AAEAAEQFRP AFSQDQGNVA 420 VEEPITTLPD LLSSETQSTD RHPYCVVDDK FDLGMQGYLD MAQGMLIDPP PIEGSSGNGG 480 DDDDGGEVKL WSY |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 164 | 174 | RERERERERAR |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00644 | PBM | Transfer from LOC_Os02g45450 | Download |
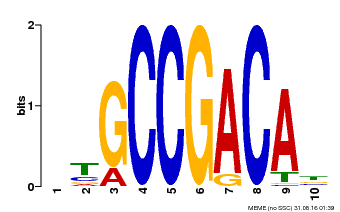
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Zpz_sc01411.1.g00210.1.am.mk |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB627353 | 0.0 | AB627353.1 Zoysia japonica gene for CBF, complete cds. | |||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP196 | 36 | 261 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G25470.1 | 9e-34 | C-repeat/DRE binding factor 2 | ||||



