 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zpz_sc01162.1.g00080.1.sm.mkhc | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 719aa MW: 78458.7 Da PI: 6.6162 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 49.1 | 1.3e-15 | 24 | 68 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r rWT+ E++++++a k++G W +I +++g ++t+ q++s+ qk+
Zpz_sc01162.1.g00080.1.sm.mkhc 24 RERWTEAEHKRFLEALKLYGRQ-WQRIEEHVG-TKTAVQIRSHAQKF 68
78******************77.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.47E-16 | 18 | 74 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 19.824 | 19 | 73 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 2.2E-16 | 22 | 71 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 1.6E-12 | 23 | 71 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 5.8E-13 | 24 | 67 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.1E-8 | 24 | 64 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.39E-9 | 26 | 69 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0042754 | Biological Process | negative regulation of circadian rhythm | ||||
| GO:0043433 | Biological Process | negative regulation of sequence-specific DNA binding transcription factor activity | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048574 | Biological Process | long-day photoperiodism, flowering | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 719 aa Download sequence Send to blast |
MEVNSSGEET VVKVRKPYTI TKQRERWTEA EHKRFLEALK LYGRQWQRIE EHVGTKTAVQ 60 IRSHAQKFFT KLEKEAMNND TSPGKAHDID IPPPRPKRKP NNPYPRKNGL NSETSTKEVS 120 NDKLTKSNIR LSSGNVEMAS HSSLQKLQRK EASEKGSCSE VLNLFREVQS ASFCSVNKTY 180 SNHGAPSGIE QAKTEIRDMT TLGKNPLSID AHEGAKEING SEMEKSNGIL ISSKSDHPHE 240 DYLDTSTQEM KLKPMSVDTA YVGQQTAKSS HSVTEKNGAT SVLATGTDGS HADQTSDHDG 300 ANGRMNPCIH PTFSVDPKFN INAAPLPCPH NYTALPPMMQ CNCNQDAYRS FINMSSTLSS 360 MLVSTLLSNP AIHAAARLAA SYWPAAENNT SVDPNQENPT ECVQGSNIGS PPSMASMVAA 420 TVAAASAWWA TQGLLPFFTP PMAFPFAPAP STAFPTTDVR ASEKGRDCQV ENTQQECPKA 480 QNQDQSESLR VAASSKSDRS GKGDLSLQTE LKISPVQIAD TTPTTVADTI DAFRNKKKQD 540 RSSCGSNTPS SSDVEGDNVP GKQDKGNDNA KQASCSNSSA GDTNNRRFRS SGSTSDSWKE 600 VSEEGRLAFD ALFSREKLPQ SFSPPQGGES KEISKEEEDE VTTVTVDLNK NAIAFDHDLD 660 TMDESRASFP NELSQLKLKS RSTGFKPYKR CSVEAKDNRV PASDDVGTKR IRLESEAST |
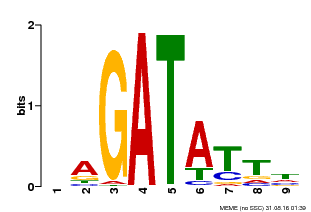
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00654 | PBM | Transfer from LOC_Os08g06110 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Zpz_sc01162.1.g00080.1.sm.mkhc |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025820794.1 | 0.0 | protein LHY isoform X1 | ||||
| Refseq | XP_025820795.1 | 0.0 | protein LHY isoform X1 | ||||
| TrEMBL | A0A2T8IF63 | 0.0 | A0A2T8IF63_9POAL; Uncharacterized protein | ||||
| STRING | Sb07g003870.1 | 0.0 | (Sorghum bicolor) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6031 | 34 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G46830.1 | 8e-39 | circadian clock associated 1 | ||||



