 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zpz_sc00046.1.g00130.1.am.mk | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 601aa MW: 65175.1 Da PI: 9.6412 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 128 | 3.7e-40 | 377 | 455 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EE.ETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvl.vsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cq+egC+adls+ak+yhrrhkvCe+hskapvv+ + gl+qrfCqqCsrfh l+efD++k+sCr+rLa+hn+rrrk+++
Zpz_sc00046.1.g00130.1.am.mk 377 RCQAEGCKADLSSAKRYHRRHKVCEHHSKAPVVVtAGGLHQRFCQQCSRFHLLDEFDDAKKSCRKRLADHNRRRRKSKP 455
6******************************98725689*************************************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 1.8E-33 | 372 | 440 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 30.291 | 375 | 453 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.27E-36 | 376 | 456 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 1.5E-30 | 378 | 452 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009554 | Biological Process | megasporogenesis | ||||
| GO:0009556 | Biological Process | microsporogenesis | ||||
| GO:0048653 | Biological Process | anther development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 601 aa Download sequence Send to blast |
MLGPEEVESG LSWLKWKSRL VANELPELES IISEMKDKMK ARHQRGVGRL ILAPLGAVDG 60 SRRARTGRAG SRSGTVVRLA APAAALIALG RNTIDRVLGH EITSHEWDAH RPSWSLPASR 120 RWRTPGAVAP SSAPAMERAA YAAAPDPAVS SSLSSFLQGT HATVPYATLK RLRYWTSQPK 180 CLSSPAASEL ASTRRERDDS VLAASSPPKK KKRKMMNIPA AGANACDEFA YVSPGAAPPS 240 LLPIMEQESS IQREQLGYNL EANSLALLPP STAAAAHQQA TIAAHDILQF YPAAAASSHY 300 LGGGGGNPYS HHFGGSNYQA YYQQAAQASP EYYFPTLVSS AEENMASFAA TQLGLNLGYR 360 TYFPPRGGYT YGHHPPRCQA EGCKADLSSA KRYHRRHKVC EHHSKAPVVV TAGGLHQRFC 420 QQCSRFHLLD EFDDAKKSCR KRLADHNRRR RKSKPTDADA ADKKRAQASK AAAGAKDKAG 480 SSSKNNMDIG DGLTTQILGS ALLSKEQDQA MDLGEVVKEA VDPKGKASMQ QQHAHHHGMS 540 NHSFPFPTSS GSCFPQSQAV SSDNTSNIAQ VQEPSLAFHQ HHQHNNILQL GQAMFDMDFD 600 H |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 1e-26 | 378 | 452 | 11 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 208 | 213 | KKKKRK |
| 2 | 447 | 464 | RRRRKSKPTDADAADKKR |
| 3 | 448 | 453 | RRRKSK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that plays an important role in building the laminar joint between leaf blade and leaf sheath boundary, thereby controlling ligule and auricle development. {ECO:0000250}. | |||||
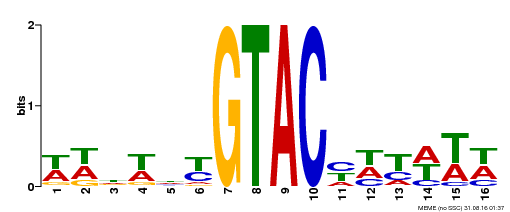
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00090 | SELEX | Transfer from AT1G02065 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Zpz_sc00046.1.g00130.1.am.mk |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | ZMU89496 | 0.0 | U89496.1 Zea mays liguleless1 protein (liguleless1) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004960135.1 | 0.0 | protein LIGULELESS 1 | ||||
| Swissprot | Q01KM7 | 0.0 | SPL8_ORYSI; Squamosa promoter-binding-like protein 8 | ||||
| TrEMBL | A0A0A9DKR3 | 0.0 | A0A0A9DKR3_ARUDO; Lg1 | ||||
| STRING | Si022315m | 0.0 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6990 | 34 | 52 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G02065.1 | 1e-40 | squamosa promoter binding protein-like 8 | ||||



