 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zmw_sc09819.1.g00010.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 287aa MW: 30108.2 Da PI: 7.9801 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 102.3 | 2.9e-32 | 109 | 162 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
pr+rWt+ LH+rFv+ave LGG+e+AtPk++lelm+vk+Ltl+hvkSHLQ+YR+
Zmw_sc09819.1.g00010.1.am.mk 109 PRMRWTTALHARFVHAVELLGGHERATPKSVLELMNVKDLTLAHVKSHLQMYRT 162
9****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 6.09E-16 | 107 | 163 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 3.8E-29 | 107 | 163 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 8.3E-24 | 109 | 162 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 3.8E-8 | 110 | 161 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005618 | Cellular Component | cell wall | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 287 aa Download sequence Send to blast |
MLRRNLPNLS LQISPPASSS DASTVAKPAP VEASTATYDD EGSGEVWLVF ANPSPSVEPP 60 DLSLELGTAA RTDDAGRRSH LLQPHHGCAF KRAAERPSVP GLKRSARAPR MRWTTALHAR 120 FVHAVELLGG HERATPKSVL ELMNVKDLTL AHVKSHLQMY RTVKSTDRSS HVTTGEALLE 180 QQRTATGMME VAVEAAAAAA EGGGRGVAAV LQRCDDMVGI CGPPAAAAAA ASAATTASAA 240 HFLCAPAPTP LMHPSSLFTC VYSSSRVGTN QGMHVPTLQS SAPEPFL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 5e-17 | 110 | 164 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 5e-17 | 110 | 164 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 5e-17 | 110 | 164 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 5e-17 | 110 | 164 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 5e-17 | 110 | 164 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 5e-17 | 110 | 164 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 5e-17 | 110 | 164 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 5e-17 | 110 | 164 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00107 | PBM | Transfer from AT5G42630 | Download |
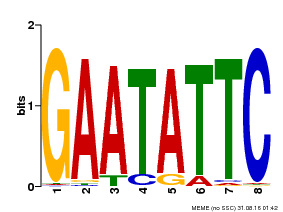
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002466395.1 | 1e-82 | probable transcription factor GLK2 | ||||
| TrEMBL | C5WZ98 | 3e-81 | C5WZ98_SORBI; Uncharacterized protein | ||||
| STRING | Sb01g007030.1 | 5e-82 | (Sorghum bicolor) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6554 | 37 | 50 |



