 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zmw_sc05144.1.g00030.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | MIKC_MADS | ||||||||
| Protein Properties | Length: 217aa MW: 24692.5 Da PI: 9.7497 | ||||||||
| Description | MIKC_MADS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SRF-TF | 94.3 | 5.7e-30 | 9 | 59 | 1 | 51 |
S---SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-TTSEEEEEE- CS
SRF-TF 1 krienksnrqvtfskRrngilKKAeELSvLCdaevaviifsstgklyeyss 51
krien + rqvtfskRrng+lKKA+ELSvLCdaeva+i+fs++gklye++s
Zmw_sc05144.1.g00030.1.sm.mkhc 9 KRIENPTSRQVTFSKRRNGLLKKAFELSVLCDAEVALIVFSTRGKLYEFAS 59
79***********************************************86 PP
| |||||||
| 2 | K-box | 50.7 | 7.7e-18 | 78 | 162 | 6 | 90 |
K-box 6 gksleeakaeslqqelakLkkeienLqreqRhllGedLesLslkeLqqLeqqLekslkkiRskKnellleqieelqkke 84
++++ +++ e+ + +++ L k++e L+ +R+llGe Le++s+ eL++Le +Leksl +R +K + l+e+ +l+k +
Zmw_sc05144.1.g00030.1.sm.mkhc 78 TNNTVQQDIEQIKADAEGLAKKLEALEAYKRKLLGERLEECSIDELHSLEVKLEKSLHIVRGRKVRKLKEEEMTLRKNN 156
44467888999******************************************************************** PP
K-box 85 kelqee 90
++l+e+
Zmw_sc05144.1.g00030.1.sm.mkhc 157 EDLREK 162
*99986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50066 | 31.819 | 1 | 61 | IPR002100 | Transcription factor, MADS-box |
| SMART | SM00432 | 1.6E-39 | 1 | 60 | IPR002100 | Transcription factor, MADS-box |
| SuperFamily | SSF55455 | 4.19E-33 | 3 | 84 | IPR002100 | Transcription factor, MADS-box |
| PROSITE pattern | PS00350 | 0 | 3 | 57 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 3.2E-31 | 3 | 23 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00265 | 3.08E-42 | 3 | 75 | No hit | No description |
| Pfam | PF00319 | 1.7E-26 | 10 | 57 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 3.2E-31 | 23 | 38 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 3.2E-31 | 38 | 59 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF01486 | 3.8E-17 | 83 | 163 | IPR002487 | Transcription factor, K-box |
| PROSITE profile | PS51297 | 11.487 | 86 | 169 | IPR002487 | Transcription factor, K-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000060 | Biological Process | protein import into nucleus, translocation | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009838 | Biological Process | abscission | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010077 | Biological Process | maintenance of inflorescence meristem identity | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0080187 | Biological Process | floral organ senescence | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005737 | Cellular Component | cytoplasm | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008134 | Molecular Function | transcription factor binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 217 aa Download sequence Send to blast |
MVRGKTQMKR IENPTSRQVT FSKRRNGLLK KAFELSVLCD AEVALIVFST RGKLYEFASA 60 SVQKTVERYR TYTKDNVTNN TVQQDIEQIK ADAEGLAKKL EALEAYKRKL LGERLEECSI 120 DELHSLEVKL EKSLHIVRGR KVRKLKEEEM TLRKNNEDLR EKCKVQPQLV APLGVVATQD 180 KHPEKAGNDM DVETELFIGL PGRDNRPSKA AIAVVEI |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5f28_A | 1e-19 | 1 | 72 | 1 | 72 | MEF2C |
| 5f28_B | 1e-19 | 1 | 72 | 1 | 72 | MEF2C |
| 5f28_C | 1e-19 | 1 | 72 | 1 | 72 | MEF2C |
| 5f28_D | 1e-19 | 1 | 72 | 1 | 72 | MEF2C |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor active in flowering time control. May control internode elongation and promote floral transition phase. May act upstream of the floral regulators MADS1, MADS14, MADS15 and MADS18 in the floral induction pathway. {ECO:0000269|PubMed:15144377, ECO:0000269|PubMed:17166135}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
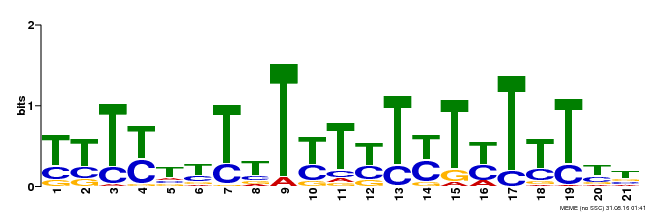
| Motif ID | Method | Source | Motif file |
| MP00076 | ChIP-chip | Transfer from AT2G45660 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004985919.1 | 1e-120 | MADS-box transcription factor 50 isoform X2 | ||||
| Refseq | XP_004985920.1 | 1e-120 | MADS-box transcription factor 50 isoform X2 | ||||
| Refseq | XP_012698462.1 | 1e-120 | MADS-box transcription factor 50 isoform X2 | ||||
| Swissprot | Q9XJ60 | 1e-112 | MAD50_ORYSJ; MADS-box transcription factor 50 | ||||
| TrEMBL | Q9FR85 | 1e-114 | Q9FR85_MAIZE; M5 protein | ||||
| STRING | GRMZM2G171365_P01 | 1e-114 | (Zea mays) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1413 | 33 | 79 |



