 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zmw_sc03239.1.g00050.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | WRKY | ||||||||
| Protein Properties | Length: 316aa MW: 34455.7 Da PI: 8.2339 | ||||||||
| Description | WRKY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | WRKY | 102.4 | 2.5e-32 | 166 | 223 | 2 | 59 |
--SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 2 dDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhe 59
+Dgy+WrKYGqK vk+s++prsYYrCt+++C vkk+vers +dp++v++tYeg+H+h+
Zmw_sc03239.1.g00050.1.sm.mkhc 166 EDGYRWRKYGQKAVKNSPYPRSYYRCTTPKCGVKKRVERSYQDPSTVITTYEGQHTHH 223
8********************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.20.25.80 | 4.4E-34 | 150 | 225 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 5.89E-29 | 158 | 225 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 31.066 | 160 | 225 | IPR003657 | WRKY domain |
| SMART | SM00774 | 1.6E-37 | 165 | 224 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 1.7E-25 | 166 | 223 | IPR003657 | WRKY domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 316 aa Download sequence Send to blast |
MESFFFSQQL GASVVGGGGR TGAEELVPPY TSITDYLQGF LDPAGLARHL DVPLSSEEVP 60 VKNEMSVDAS LDSQGTSGVA GDGAAMLTPN SSGSLSSSDR EGEGQPRRSK KGIRAKQAED 120 ADAVDQKDLE EGENSMKANK PKKKAEKRQR QPRVAFLTKS EVDHLEDGYR WRKYGQKAVK 180 NSPYPRSYYR CTTPKCGVKK RVERSYQDPS TVITTYEGQH THHSPASLRA GGGHLFMPNA 240 HTLPPHLMPS AGFRADLMSM MHPISMGANP NMYLPSMPPP PPPPVPTSPA PPLHQSHFTD 300 YALLQDLFPS NMPNNL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wj2_A | 5e-30 | 158 | 226 | 10 | 78 | Probable WRKY transcription factor 4 |
| 2lex_A | 5e-30 | 158 | 226 | 10 | 78 | Probable WRKY transcription factor 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. Interacts specifically with the W box (5'-(T)TGAC[CT]-3'), a frequently occurring elicitor-responsive cis-acting element (By similarity). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00440 | DAP | Transfer from AT4G18170 | Download |
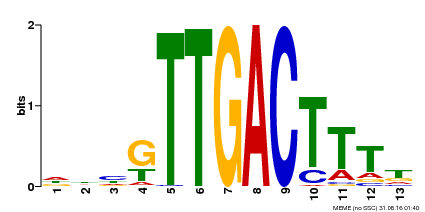
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001147623.1 | 1e-154 | uncharacterized protein LOC100281232 | ||||
| Swissprot | Q8VWJ2 | 5e-48 | WRK28_ARATH; WRKY transcription factor 28 | ||||
| TrEMBL | A0A3L6F9V2 | 1e-156 | A0A3L6F9V2_MAIZE; Putative WRKY transcription factor 71 | ||||
| STRING | AC198725.4_FGP009 | 1e-154 | (Zea mays) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP623 | 37 | 173 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



