 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zmw_sc02154.1.g00030.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 482aa MW: 51604.7 Da PI: 8.3746 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 56.6 | 6.4e-18 | 310 | 360 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
+ ykGVr++ grWv+e+r+p g ++r++lg+f tae+Aa+a++ a+++l+g
Zmw_sc02154.1.g00030.1.am.mk 310 PVYKGVRRRN-PGRWVCEVREP--HG-KQRIWLGTFETAEMAARAHDVAALALRG 360
68*****887.9******9996..33.5*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF00847 | 2.6E-13 | 310 | 360 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 3.4E-29 | 311 | 370 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 1.4E-26 | 311 | 374 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 21.694 | 311 | 368 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 1.77E-20 | 311 | 369 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 8.83E-27 | 312 | 370 | No hit | No description |
| PRINTS | PR00367 | 9.4E-8 | 312 | 323 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 9.4E-8 | 334 | 350 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009631 | Biological Process | cold acclimation | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 482 aa Download sequence Send to blast |
MMRAMPKTLV GGGKLGKAES ARDGAEDRDG QPNSVDIAPK FSDDRVENNS NSVAGARLWL 60 HPLAAADLGA TCDVNRQTNG PVYPSGHGES PSGPSPLAID LEPQRPGPMK LPLNRDAGRI 120 LCAALDASFA LVRRVRRAED PIPAITAVAE ERERERERER ERERAPGWGE AQAGGRRPMT 180 PRCARPNRSS APRAPAGSDP TNLRSPLARE PVPRVLDPSS RANLAGISGA ATRRSSAHGH 240 VALDAPSTLP SRRSIRSIAM DVGALSSDYS SGTPSPVGGS DGSSASYMTV SSAPPKRRAG 300 RTKFKETRHP VYKGVRRRNP GRWVCEVREP HGKQRIWLGT FETAEMAARA HDVAALALRG 360 LGACLNFADS PRLLRVPPLG AGHDEIRRAA AEAAEQFRPA FSQDQGNVAV EEPITTLPDL 420 LSSETQSTDR HPYCVVDDKF DLGMQGYLDM AQGMLIDPPP IEGSSGNGGD DDDGGEVKLW 480 SY |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 153 | 163 | RERERERERER |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00644 | PBM | Transfer from LOC_Os02g45450 | Download |
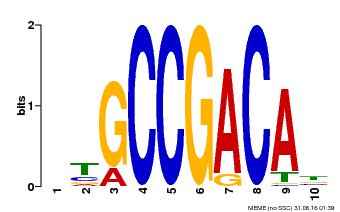
| |||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP196 | 36 | 261 |



