 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zmw_sc01917.1.g00130.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 560aa MW: 61097 Da PI: 7.2899 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 175.8 | 1.2e-54 | 143 | 272 | 1 | 129 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLpk...kvkaeekewyfFskrdkkyatgkrknrat 78
+ppGfrFhPtdeelv +yL+kkv+++k++l +vi+++d+y++ePwdL++ + e++ewyfFs +d+ky+tg+r+nrat
Zmw_sc01917.1.g00130.1.am.mk 143 VPPGFRFHPTDEELVGYYLRKKVASQKIDL-DVIRDIDLYRIEPWDLQEhcgIGYDEQNEWYFFSYKDRKYPTGTRTNRAT 222
69****************************.9***************953432234677********************** PP
NAM 79 ksgyWkatgkdkevlskkgelvglkktLvfykgrapkgektdWvmheyrle 129
+g+Wkatg+dk+v++ k++l+g++ktLvfykgrap+g+ktdW+mheyrle
Zmw_sc01917.1.g00130.1.am.mk 223 MAGFWKATGRDKAVHD-KSRLIGMRKTLVFYKGRAPNGQKTDWIMHEYRLE 272
****************.999*****************************85 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 2.22E-58 | 140 | 311 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 57.856 | 143 | 311 | IPR003441 | NAC domain |
| Pfam | PF02365 | 3.5E-28 | 144 | 271 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0048759 | Biological Process | xylem vessel member cell differentiation | ||||
| GO:1901348 | Biological Process | positive regulation of secondary cell wall biogenesis | ||||
| GO:1990110 | Biological Process | callus formation | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 560 aa Download sequence Send to blast |
MEFLDQATER SEAVQATETH LLLVLPGFSL AILPRPVHAR HAFVAPLDGG DGGPEVAFPL 60 LPAAPPALSK VLNYHARRPA CMQLSKVVQA RALPPPAARQ KLANAISFCY LHYLQGSGVA 120 ESDAHTYSGA APGDHMDSME SCVPPGFRFH PTDEELVGYY LRKKVASQKI DLDVIRDIDL 180 YRIEPWDLQE HCGIGYDEQN EWYFFSYKDR KYPTGTRTNR ATMAGFWKAT GRDKAVHDKS 240 RLIGMRKTLV FYKGRAPNGQ KTDWIMHEYR LETDENAPPQ ARPLAPLSFT PNKLVVVLQE 300 EGWVVCRAFK KRTAYPARSM ALAWDSSFAY RDPNVMVGAT AAEAAAFVDP NKAYAQIRRQ 360 SSKNARFKQE AELDGAAAFL QYSSHLVELP QLESPSAPLA PNTSHASTEE GGSGGGRRSK 420 KARADKVATD WRALDKFVAS QLSPGEGGDI EATASAAAAG EGSQLDHGCE EDMAALLFLN 480 SDGREEVERW TGLLGPASGR WPPLPTRVGL GMHGHGLRPT YVHGPCSHTP VATCDPSLLT 540 NFVVGPSPAG GAADLRRGRD |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 3ulx_A | 6e-47 | 140 | 315 | 12 | 172 | Stress-induced transcription factor NAC1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the secondary wall NAC binding element (SNBE), 5'-(T/A)NN(C/T)(T/C/G)TNNNNNNNA(A/C)GN(A/C/T)(A/T)-3', in the promoter of target genes (By similarity). Involved in xylem formation by promoting the expression of secondary wall-associated transcription factors and of genes involved in secondary wall biosynthesis and programmed cell death, genes driven by the secondary wall NAC binding element (SNBE). Triggers thickening of secondary walls (PubMed:25148240). {ECO:0000250|UniProtKB:Q9LVA1, ECO:0000269|PubMed:25148240}. | |||||
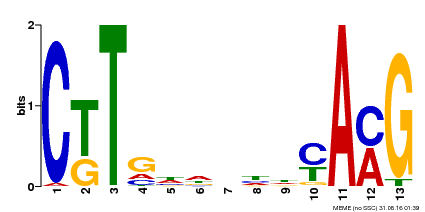
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00266 | DAP | Transfer from AT2G18060 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by chitin (e.g. chitooctaose) (PubMed:17722694). Accumulates in plants exposed to callus induction medium (CIM) (PubMed:17581762). {ECO:0000269|PubMed:17581762, ECO:0000269|PubMed:17722694}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_022678630.1 | 0.0 | NAC domain-containing protein 37 isoform X1 | ||||
| Swissprot | Q9SL41 | 1e-116 | NAC37_ARATH; NAC domain-containing protein 37 | ||||
| TrEMBL | A0A368SWG2 | 0.0 | A0A368SWG2_SETIT; Uncharacterized protein | ||||
| STRING | Pavir.Ib00161.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3625 | 37 | 68 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



