 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zmw_sc01681.1.g00090.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 267aa MW: 29182.3 Da PI: 4.5118 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 61.6 | 1.2e-19 | 45 | 98 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k++++++eq+++Le+ Fe ++++ e++++LA+ lgL+ rqV vWFqNrRa++k
Zmw_sc01681.1.g00090.1.am.mk 45 KKRRLSTEQVRALERSFEVENKLEPERKARLARDLGLQPRQVAVWFQNRRARWK 98
45689************************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 135.1 | 2.4e-43 | 45 | 136 | 2 | 93 |
HD-ZIP_I/II 2 kkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLek 82
kkrrls+eqv++LE+sFe e+kLeperK++lar+LglqprqvavWFqnrRAR+ktkqlE+dy+aL+++yd+l++++++L++
Zmw_sc01681.1.g00090.1.am.mk 45 KKRRLSTEQVRALERSFEVENKLEPERKARLARDLGLQPRQVAVWFQNRRARWKTKQLERDYAALRHSYDTLRHDHDALRR 125
9******************************************************************************** PP
HD-ZIP_I/II 83 eveeLreelke 93
++++L +e+ke
Zmw_sc01681.1.g00090.1.am.mk 126 DKDALLAEIKE 136
******99987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 5.56E-20 | 35 | 102 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 2.7E-18 | 43 | 104 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.00E-18 | 45 | 101 | No hit | No description |
| PROSITE profile | PS50071 | 17.135 | 45 | 100 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 6.9E-17 | 45 | 98 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 2.6E-20 | 47 | 107 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 9.0E-6 | 71 | 80 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 75 | 98 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 9.0E-6 | 80 | 96 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 2.8E-17 | 100 | 141 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009637 | Biological Process | response to blue light | ||||
| GO:0030308 | Biological Process | negative regulation of cell growth | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048510 | Biological Process | regulation of timing of transition from vegetative to reproductive phase | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 267 aa Download sequence Send to blast |
MKRSRGGSPS LLSMTNPDDD GYDGVGMEAE GDAEEEMMVC GGGGKKRRLS TEQVRALERS 60 FEVENKLEPE RKARLARDLG LQPRQVAVWF QNRRARWKTK QLERDYAALR HSYDTLRHDH 120 DALRRDKDAL LAEIKELKAK LGDEEAAASF TSVKAEPATS DGPPPTGAGS SESDSSAVLN 180 DADQAGATPP TEAPVPEIQP GTLVGTATVM AGTAPARQGE VFFHGSFLKA EEDETGFLDD 240 DEPCGGFFAV EQPPPMAWWT EPIEHWN |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 69 | 75 | ERKARLA |
| 2 | 92 | 100 | RRARWKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription activator that binds to the DNA sequence 5'-CAAT[AT]ATTG-3'. May be involved in the regulation of gibberellin signaling. {ECO:0000269|PubMed:10732669, ECO:0000269|PubMed:18049796}. | |||||
| UniProt | Probable transcription activator that binds to the DNA sequence 5'-CAAT[AT]ATTG-3'. May be involved in the regulation of gibberellin signaling. {ECO:0000269|PubMed:10732669, ECO:0000269|PubMed:18049796}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00051 | PBM | Transfer from AT4G40060 | Download |
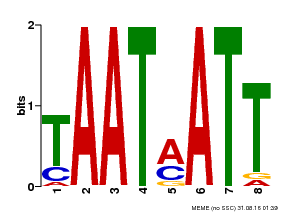
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By gibberellin. Down-regulated in leaves by drought stress. {ECO:0000269|PubMed:17999151, ECO:0000269|PubMed:18049796}. | |||||
| UniProt | INDUCTION: By gibberellin. Down-regulated in leaves by drought stress. {ECO:0000269|PubMed:17999151, ECO:0000269|PubMed:18049796}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004957072.1 | 1e-153 | homeobox-leucine zipper protein HOX4 | ||||
| Swissprot | Q6K498 | 1e-128 | HOX4_ORYSJ; Homeobox-leucine zipper protein HOX4 | ||||
| Swissprot | Q9XH37 | 1e-128 | HOX4_ORYSI; Homeobox-leucine zipper protein HOX4 | ||||
| TrEMBL | K3ZW40 | 1e-152 | K3ZW40_SETIT; Uncharacterized protein | ||||
| STRING | Si030822m | 1e-153 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3009 | 34 | 82 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



