 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zmw_sc00963.1.g00120.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 334aa MW: 36815.1 Da PI: 4.1791 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 63.8 | 2.5e-20 | 75 | 128 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k++++t+eq++ Le+ Fe+++++ e+++eLA+klg++ rqV vWFqNrRa++k
Zmw_sc00963.1.g00120.1.am.mkhc 75 KKRRLTPEQVHLLERSFEEENKLEPERKTELARKLGMQPRQVAVWFQNRRARWK 128
5678*************************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 128.3 | 3.2e-41 | 74 | 165 | 1 | 92 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeener 79
ekkrrl+ eqv+lLE+sFeee+kLeperK+elar+Lg+qprqvavWFqnrRAR+ktkqlE+d+++Lk++++al++e +
Zmw_sc00963.1.g00120.1.am.mkhc 74 EKKRRLTPEQVHLLERSFEEENKLEPERKTELARKLGMQPRQVAVWFQNRRARWKTKQLEQDFDRLKASFNALRAEYDV 152
69***************************************************************************** PP
HD-ZIP_I/II 80 LekeveeLreelk 92
L +++++Lr+++
Zmw_sc00963.1.g00120.1.am.mkhc 153 LLQDNHRLRTQVV 165
*********9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 3.22E-20 | 68 | 132 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.912 | 70 | 130 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.9E-20 | 73 | 134 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 1.4E-17 | 75 | 128 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 6.15E-20 | 75 | 131 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 7.0E-22 | 76 | 137 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 4.5E-6 | 101 | 110 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 105 | 128 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 4.5E-6 | 110 | 126 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 1.3E-15 | 130 | 172 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0001558 | Biological Process | regulation of cell growth | ||||
| GO:0009637 | Biological Process | response to blue light | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 334 aa Download sequence Send to blast |
MDPGRLIFNT TGSGAGQMLF LDCGVGGPAG GMFHRGGRPV FAGGLEDGRG VKRPFFTSPD 60 ELLEEEYYDE QLPEKKRRLT PEQVHLLERS FEEENKLEPE RKTELARKLG MQPRQVAVWF 120 QNRRARWKTK QLEQDFDRLK ASFNALRAEY DVLLQDNHRL RTQVVSLTEK LQEKEDAESG 180 ASATADAAEI TTDAKQDSLA LDAQETAAEV AFDAQQVKSE DRLSTGSDES AVGDTDALLG 240 AGGGLGTVDS SFESYFPGGD DHYNDCGMEP VDHCAAEFQS EEDDGAGSDE GCSYYADETA 300 AAVAFFVGAH AHHQVQPVDD DEDDGQISSW WMWN |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 122 | 130 | RRARWKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
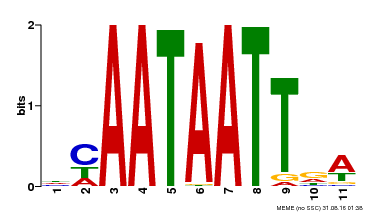
| Motif ID | Method | Source | Motif file |
| MP00323 | DAP | Transfer from AT3G01470 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025813024.1 | 1e-142 | homeobox-leucine zipper protein HOX16-like isoform X1 | ||||
| Swissprot | Q6YWR4 | 1e-130 | HOX16_ORYSJ; Homeobox-leucine zipper protein HOX16 | ||||
| TrEMBL | A0A2T7FC04 | 1e-143 | A0A2T7FC04_9POAL; Uncharacterized protein | ||||
| STRING | Pavir.Aa00462.1.p | 1e-143 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2609 | 38 | 87 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



