 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zosma76g00390.1 | ||||||||
| Common Name | ZOSMA_76G00390 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Alismatales; Zosteraceae; Zostera
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 717aa MW: 79177.5 Da PI: 7.9416 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 45.8 | 1.4e-14 | 37 | 81 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r rWT+eE+ ++++a k++G W I +++ ++t+ q++s+ qk+
Zosma76g00390.1 37 RERWTEEEHNRFLQALKLYGRA-WQHIQEYVA-TKTAVQIRSHAQKF 81
78******************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 3.51E-16 | 31 | 87 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 19.985 | 32 | 86 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 6.2E-16 | 35 | 84 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 1.5E-12 | 36 | 84 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.5E-12 | 37 | 80 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.3E-8 | 37 | 77 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 4.03E-9 | 39 | 82 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010243 | Biological Process | response to organonitrogen compound | ||||
| GO:0042754 | Biological Process | negative regulation of circadian rhythm | ||||
| GO:0043496 | Biological Process | regulation of protein homodimerization activity | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048574 | Biological Process | long-day photoperiodism, flowering | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 717 aa Download sequence Send to blast |
MMFIDLNHLN LTMDEASSFG EDFNVKARKP YTITKQRERW TEEEHNRFLQ ALKLYGRAWQ 60 HIQEYVATKT AVQIRSHAQK FFSKLEKEAL SKGVVPGQTI NIDIPPPRPK RKPTNPYPRK 120 TCMGTISSSK EGNDKTSSIF SSCQNKTQLS NGLSLENPAR SESSEGTEQA SENCDCSEVF 180 TLFHNDPCAS CTSSVKNCGN PSSIKKFIPI LEDINHTSEM IKDRNSQIAI DNKEAMQYTD 240 NDCDVDIGRL EGIKKSNLNT NSASMEKKAK HKKPDEATSN CLMQDTNERS KKQKCCVQSN 300 SQISQSDIHV CHIPELINES MPAKPEHHTK STTSFNQEHP SSPPFTFFHG NQDVYRSLIN 360 MSSSFSNCIL FCLQQNPSAH VAANLAASVW HCLEGDGMNS TAKVVDGGNY PNNAYTAPNM 420 VAIASATVAA ATAWWATHGL HTFPHPHSTD LNFAGVAPII TNPVSEGREV RKDDSVQFQA 480 QQDERILDSE SPQTAVLQCS TSKSTSSSDC GESKRGDEVI GGNTTKSDRK VNCYSYGSSK 540 LSSTEKYTNV LLKNTESQTL EQVMKSNKLE NQTILSEANG QRIKGSMVVN DSWKEVSEEG 600 RLAFEALFSR EVLPQSFSLT GLTAMGMTPR NLKMGNVKAG DAKRKVSADV VADNDVDMGN 660 SRMKCKLNGN LNLERSGFKP YKRCSMDAKK GTTSQGQGDE KGSRKRLRLE GETTAR* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00654 | PBM | Transfer from LOC_Os08g06110 | Download |
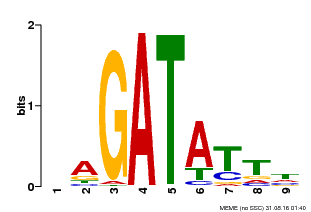
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| TrEMBL | A0A0K9NPD1 | 0.0 | A0A0K9NPD1_ZOSMR; LATE ELONGATED HYPOCOTYL transcription factor | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6031 | 34 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G46830.1 | 1e-62 | circadian clock associated 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Zosma76g00390.1 |



