 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zosma54g00840.1 | ||||||||
| Common Name | ZOSMA_54G00840 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Alismatales; Zosteraceae; Zostera
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 297aa MW: 32808.2 Da PI: 4.1818 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 62.1 | 8.6e-20 | 81 | 135 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakekk 57
k++++t+eq++ Le+ Fe + ++ e++ eL+kklgL+ rqV vWFqNrRa++k+
Zosma54g00840.1 81 KKRRLTPEQVHMLETSFETEIKLEPERKMELSKKLGLQPRQVAVWFQNRRARWKN 135
5678*************************************************95 PP
| |||||||
| 2 | HD-ZIP_I/II | 121.1 | 5.4e-39 | 80 | 168 | 1 | 89 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLre 89
ekkrrl+ eqv++LE+sFe+e kLeperK+el+++LglqprqvavWFqnrRAR+k+kqlE+++ +Lk+ yd+l+++++ L +++++L +
Zosma54g00840.1 80 EKKRRLTPEQVHMLETSFETEIKLEPERKMELSKKLGLQPRQVAVWFQNRRARWKNKQLETEFCQLKSLYDTLRANHDCLLQNNDSLLA 168
69***************************************************************************999999998864 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 8.98E-20 | 74 | 138 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 16.876 | 76 | 136 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 3.5E-19 | 79 | 140 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 7.97E-19 | 81 | 137 | No hit | No description |
| Pfam | PF00046 | 5.0E-17 | 81 | 135 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 1.7E-20 | 82 | 140 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 1.0E-5 | 107 | 116 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 111 | 134 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 1.0E-5 | 116 | 132 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 1.4E-7 | 136 | 168 | IPR003106 | Leucine zipper, homeobox-associated |
| Pfam | PF02183 | 3.7E-10 | 149 | 185 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0001558 | Biological Process | regulation of cell growth | ||||
| GO:0009637 | Biological Process | response to blue light | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 297 aa Download sequence Send to blast |
MDSSRFGFEY SASGPDNGNV FFVGRSAVLG FPDVGGGTSL STEDVSSGVG GSKRRTSIFS 60 SSEELIYGDD LYYDEQLLPE KKRRLTPEQV HMLETSFETE IKLEPERKME LSKKLGLQPR 120 QVAVWFQNRR ARWKNKQLET EFCQLKSLYD TLRANHDCLL QNNDSLLADN HRLRSEVISL 180 TGKLQEIKNS SGTSEVLDGV NGAVTLVSPL PESTDVDLDG APLVMESTDL YYTEVVEDGS 240 FSFHGAGCQF TVTCEDYDGS DDGGVYFPGA FSENPGADEL LEKDAESDII DWSVWS* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 128 | 136 | RRARWKNKQ |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
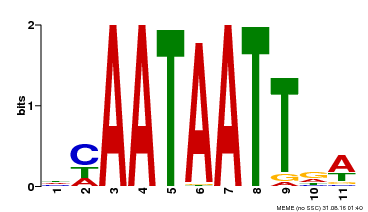
| Motif ID | Method | Source | Motif file |
| MP00323 | DAP | Transfer from AT3G01470 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| TrEMBL | A0A0K9NYV1 | 0.0 | A0A0K9NYV1_ZOSMR; Homeobox-leucine zipper protein HAT5 | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2609 | 38 | 87 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G01470.1 | 1e-48 | homeobox 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Zosma54g00840.1 |



