 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zosma291g00060.1 | ||||||||
| Common Name | ZOSMA_291G00060 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Alismatales; Zosteraceae; Zostera
|
||||||||
| Family | WRKY | ||||||||
| Protein Properties | Length: 704aa MW: 76009 Da PI: 5.768 | ||||||||
| Description | WRKY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | WRKY | 99.8 | 1.6e-31 | 357 | 413 | 2 | 59 |
--SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 2 dDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhe 59
dDgynWrKYGqK+vkgsefprsYY+Ct+ +C+vkkkver+ ++v+ei+Y+g Hnh+
Zosma291g00060.1 357 DDGYNWRKYGQKHVKGSEFPRSYYKCTHLNCQVKKKVERAP-GGQVTEIIYKGAHNHS 413
8****************************************.***************7 PP
| |||||||
| 2 | WRKY | 102.9 | 1.8e-32 | 551 | 608 | 1 | 58 |
---SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS CS
WRKY 1 ldDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnh 58
ldDgy+WrKYGqK+vkg+++prsYY+Cts gC+v+k+ver+++d k+v++tYeg+Hnh
Zosma291g00060.1 551 LDDGYRWRKYGQKVVKGNPNPRSYYKCTSVGCTVRKHVERASHDLKAVITTYEGKHNH 608
59******************************************************** PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50811 | 22.538 | 351 | 415 | IPR003657 | WRKY domain |
| Gene3D | G3DSA:2.20.25.80 | 6.7E-27 | 354 | 415 | IPR003657 | WRKY domain |
| SMART | SM00774 | 3.0E-34 | 356 | 414 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 1.14E-24 | 356 | 415 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 2.7E-24 | 357 | 412 | IPR003657 | WRKY domain |
| Gene3D | G3DSA:2.20.25.80 | 2.7E-36 | 536 | 611 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 6.54E-28 | 543 | 611 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 37.677 | 546 | 611 | IPR003657 | WRKY domain |
| SMART | SM00774 | 3.7E-37 | 551 | 610 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 1.6E-24 | 552 | 608 | IPR003657 | WRKY domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009555 | Biological Process | pollen development | ||||
| GO:0009942 | Biological Process | longitudinal axis specification | ||||
| GO:0030010 | Biological Process | establishment of cell polarity | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 704 aa Download sequence Send to blast |
MAGSDDNLPM LDWPITTPRT QAFLSSILGG DGLLPPENGD EEVPLLATVS SASEKQHNLC 60 IDSIVDREEI DGIRRIQDLG DQSTSIADTS VPNFPPKHHG FRSGLAERMA ARGGFNAPKL 120 NTYQIRSSNN LHSSQPPSDG DAKSHQIPLS PGFSPSAFLD SPVFLFNSLP QPSPTTGKIT 180 LGQGYGSNNT SSAVPGFSNN ISSAVPGFSN NTSSAVPGEP PSEVETKTIS SKPPGEPPAS 240 YFSNASNKVH HHEQSLQILE VISTEEELSN QSESIDIHLQ KQKQQDNDLL SAFSEPSDIK 300 DSSVDYIFDI KPSQFPSMMC SKENSPPPVD QLKSEDQKVE FSSNYAGASV GAKPSDDDGY 360 NWRKYGQKHV KGSEFPRSYY KCTHLNCQVK KKVERAPGGQ VTEIIYKGAH NHSRPPLNRR 420 ISSVAVMSSH SLGAEMQTDV TSSSTSFAPL ENDNFCDPSV SSMCEDIPPR FESSDTLDVS 480 SALSNGGEED NGGTMTCGGG GYGATVGVDN EESELKRRKL ELSAVDMNSS ASRAVREPRV 540 VVQTTSEVDI LDDGYRWRKY GQKVVKGNPN PRSYYKCTSV GCTVRKHVER ASHDLKAVIT 600 TYEGKHNHVV PAARNSCHSS SVISAAAATE TTNASSHKTQ ESLTRFDIPQ TGFSFGLTQP 660 NLTSFAMAGL VPCGPPHMKP HPYLDQPGLF LLPKGEPKDD RCE* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wj2_A | 2e-35 | 357 | 612 | 8 | 78 | Probable WRKY transcription factor 4 |
| 2lex_A | 2e-35 | 357 | 612 | 8 | 78 | Probable WRKY transcription factor 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. Regulates WOX8 and WOX9 expression and basal cell division patterns during early embryogenesis. Interacts specifically with the W box (5'-(T)TGAC[CT]-3'), a frequently occurring elicitor-responsive cis-acting element. Required to repolarize the zygote from a transient symmetric state. {ECO:0000269|PubMed:21316593}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00071 | PBM | Transfer from AT5G56270 | Download |
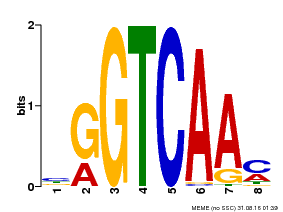
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Swissprot | Q9FG77 | 7e-94 | WRKY2_ARATH; Probable WRKY transcription factor 2 | ||||
| TrEMBL | A0A0K9PCD0 | 0.0 | A0A0K9PCD0_ZOSMR; Putative WRKY transcription factor 20 | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2075 | 38 | 100 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G56270.1 | 1e-86 | WRKY DNA-binding protein 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Zosma291g00060.1 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



