 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT5G56270.1 | ||||||||
| Common Name | ATWRKY2, K24C1.9, MXK23.1, WRKY2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | WRKY | ||||||||
| Protein Properties | Length: 687aa MW: 74561.1 Da PI: 5.9843 | ||||||||
| Description | WRKY DNA-binding protein 2 | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | WRKY | 100.1 | 1.4e-31 | 273 | 329 | 2 | 59 |
--SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 2 dDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhe 59
+DgynWrKYGqK vkgse+prsYY+Ct ++C+vkkkvers+ +++++ei+Y+g Hnh
AT5G56270.1 273 EDGYNWRKYGQKLVKGSEYPRSYYKCTNPNCQVKKKVERSR-EGHITEIIYKGAHNHL 329
8****************************************.***************5 PP
| |||||||
| 2 | WRKY | 103.8 | 9.2e-33 | 486 | 544 | 1 | 59 |
---SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 1 ldDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhe 59
ldDgy+WrKYGqK+vkg+++prsYY+Ct +gC+v+k+ver+++d k v++tYeg+Hnh+
AT5G56270.1 486 LDDGYRWRKYGQKVVKGNPNPRSYYKCTAPGCTVRKHVERASHDLKSVITTYEGKHNHD 544
59********************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.20.25.80 | 7.3E-28 | 267 | 331 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 23.169 | 267 | 331 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 1.31E-24 | 270 | 331 | IPR003657 | WRKY domain |
| SMART | SM00774 | 2.1E-34 | 272 | 330 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 5.1E-24 | 273 | 328 | IPR003657 | WRKY domain |
| Gene3D | G3DSA:2.20.25.80 | 9.9E-37 | 471 | 546 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 1.57E-28 | 478 | 546 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 38.136 | 481 | 546 | IPR003657 | WRKY domain |
| SMART | SM00774 | 1.8E-37 | 486 | 545 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 8.9E-25 | 487 | 544 | IPR003657 | WRKY domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009555 | Biological Process | pollen development | ||||
| GO:0009942 | Biological Process | longitudinal axis specification | ||||
| GO:0030010 | Biological Process | establishment of cell polarity | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000013 | anatomy | cauline leaf | ||||
| PO:0000037 | anatomy | shoot apex | ||||
| PO:0000084 | anatomy | plant sperm cell | ||||
| PO:0000230 | anatomy | inflorescence meristem | ||||
| PO:0000293 | anatomy | guard cell | ||||
| PO:0008019 | anatomy | leaf lamina base | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009006 | anatomy | shoot system | ||||
| PO:0009009 | anatomy | plant embryo | ||||
| PO:0009025 | anatomy | vascular leaf | ||||
| PO:0009029 | anatomy | stamen | ||||
| PO:0009030 | anatomy | carpel | ||||
| PO:0009031 | anatomy | sepal | ||||
| PO:0009032 | anatomy | petal | ||||
| PO:0009046 | anatomy | flower | ||||
| PO:0009047 | anatomy | stem | ||||
| PO:0020030 | anatomy | cotyledon | ||||
| PO:0020038 | anatomy | petiole | ||||
| PO:0020100 | anatomy | hypocotyl | ||||
| PO:0020137 | anatomy | leaf apex | ||||
| PO:0025022 | anatomy | collective leaf structure | ||||
| PO:0025195 | anatomy | pollen tube cell | ||||
| PO:0025281 | anatomy | pollen | ||||
| PO:0001016 | developmental stage | L mature pollen stage | ||||
| PO:0001017 | developmental stage | M germinated pollen stage | ||||
| PO:0001054 | developmental stage | vascular leaf senescent stage | ||||
| PO:0001078 | developmental stage | plant embryo cotyledonary stage | ||||
| PO:0001185 | developmental stage | plant embryo globular stage | ||||
| PO:0004507 | developmental stage | plant embryo bilateral stage | ||||
| PO:0007064 | developmental stage | LP.12 twelve leaves visible stage | ||||
| PO:0007095 | developmental stage | LP.08 eight leaves visible stage | ||||
| PO:0007098 | developmental stage | LP.02 two leaves visible stage | ||||
| PO:0007103 | developmental stage | LP.10 ten leaves visible stage | ||||
| PO:0007115 | developmental stage | LP.04 four leaves visible stage | ||||
| PO:0007123 | developmental stage | LP.06 six leaves visible stage | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| PO:0007616 | developmental stage | flowering stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 687 aa Download sequence Send to blast |
MAGFDENVAV MGEWVPRSPS PGTLFSSAIG EEKSSKRVLE RELSLNHGQV IGLEEDTSSN 60 HNKDSSQSNV FRGGLSERIA ARAGFNAPRL NTENIRTNTD FSIDSNLRSP CLTISSPGLS 120 PATLLESPVF LSNPLAQPSP TTGKFPFLPG VNGNALSSEK AKDEFFDDIG ASFSFHPVSR 180 SSSSFFQGTT EMMSVDYGNY NNRSSSHQSA EEVKPGSENI ESSNLYGIET DNQNGQNKTS 240 DVTTNTSLET VDHQEEEEEQ RRGDSMAGGA PAEDGYNWRK YGQKLVKGSE YPRSYYKCTN 300 PNCQVKKKVE RSREGHITEI IYKGAHNHLK PPPNRRSGMQ VDGTEQVEQQ QQQRDSAATW 360 VSCNNTQQQG GSNENNVEEG STRFEYGNQS GSIQAQTGGQ YESGDPVVVV DASSTFSNDE 420 DEDDRGTHGS VSLGYDGGGG GGGGEGDESE SKRRKLEAFA AEMSGSTRAI REPRVVVQTT 480 SDVDILDDGY RWRKYGQKVV KGNPNPRSYY KCTAPGCTVR KHVERASHDL KSVITTYEGK 540 HNHDVPAARN SSHGGGGDSG NGNSGGSAAV SHHYHNGHHS EPPRGRFDRQ VTTNNQSPFS 600 RPFSFQPHLG PPSGFSFGLG QTGLVNLSMP GLAYGQGKMP GLPHPYMTQP VGMSEAMMQR 660 GMEPKVEPVS DSGQSVYNQI MSRLPQI |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wj2_A | 4e-37 | 273 | 547 | 8 | 78 | Probable WRKY transcription factor 4 |
| 2lex_A | 4e-37 | 273 | 547 | 8 | 78 | Probable WRKY transcription factor 4 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| At.21527 | 0.0 | floral meristem | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 30696720 | 0.0 | ||||
| Genevisible | 248008_at | 0.0 | ||||
| Expression Atlas | AT5G56270 | - | ||||
| AtGenExpress | AT5G56270 | - | ||||
| ATTED-II | AT5G56270 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Low expression in senescent leaves (PubMed:11722756). Expressed in both the unfertilized egg cell and the pollen tube (PubMed:21316593). {ECO:0000269|PubMed:11722756, ECO:0000269|PubMed:21316593}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| TAIR | WRKY transcription factor 2 | |||||
| UniProt | Transcription factor. Regulates WOX8 and WOX9 expression and basal cell division patterns during early embryogenesis. Interacts specifically with the W box (5'-(T)TGAC[CT]-3'), a frequently occurring elicitor-responsive cis-acting element. Required to repolarize the zygote from a transient symmetric state. {ECO:0000269|PubMed:21316593}. | |||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
|
||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00071 | PBM | 25215497 | Download |
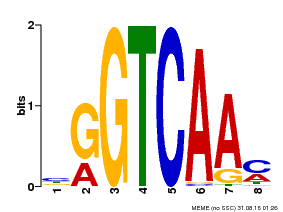
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT5G56270.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Regulation -- ATRM (Manually Curated Upstream Regulators) ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator (A: Activate/R: Repress) | |||||
| ATRM | AT2G36270 (A), AT3G24650 (A) | |||||
| Regulation -- ATRM (Manually Curated Target Genes) ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Target Gene (A: Activate/R: Repress) | |||||
| ATRM | AT2G36270(A), AT2G40170(R), AT3G24650(A), AT3G51810(R) | |||||
| Regulation -- Hormone ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hormone | |||||
| AHD | abscisic acid | |||||
| Phenotype -- Disruption Phenotype ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | DISRUPTION PHENOTYPE: Strong defects in embryo development. {ECO:0000269|PubMed:21316593}. | |||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT5G56270 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF418308 | 0.0 | AF418308.1 Arabidopsis thaliana WRKY transcription factor 2 (WRKY2) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_200438.1 | 0.0 | WRKY DNA-binding protein 2 | ||||
| Swissprot | Q9FG77 | 0.0 | WRKY2_ARATH; Probable WRKY transcription factor 2 | ||||
| TrEMBL | A0A178US56 | 0.0 | A0A178US56_ARATH; WRKY2 | ||||
| STRING | AT5G56270.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM7954 | 27 | 40 | Representative plant | OGRP14 | 17 | 875 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT5G56270.1 |
| Entrez Gene | 835726 |
| iHOP | AT5G56270 |
| wikigenes | AT5G56270 |



