 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zosma237g00270.1 | ||||||||
| Common Name | ZOSMA_237G00270 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Alismatales; Zosteraceae; Zostera
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 219aa MW: 23639.5 Da PI: 9.6951 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 54.1 | 3.8e-17 | 42 | 91 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
+y+G+r +k +g+WvAe+r+p +kr r++lg++ t+ Aaka++ a+ l+g
Zosma237g00270.1 42 KYRGIRMRK-WGKWVAEVREP---NKRSRIWLGSYSTPIAAAKAYDTAMFHLRG 91
8******99.*******9998...337************************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00380 | 9.2E-35 | 42 | 105 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 22.352 | 42 | 99 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 4.64E-20 | 42 | 101 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 5.02E-26 | 42 | 99 | No hit | No description |
| Gene3D | G3DSA:3.30.730.10 | 1.9E-29 | 42 | 99 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 4.4E-11 | 42 | 91 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.2E-11 | 43 | 54 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.2E-11 | 81 | 101 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 219 aa Download sequence Send to blast |
MEKRECCSNS LPSSSSSSPT PTTAYTASHS SETLPCGSQE IKYRGIRMRK WGKWVAEVRE 60 PNKRSRIWLG SYSTPIAAAK AYDTAMFHLR GRSARLNFPM EFADKGVDGS STGGGGVLSS 120 ETIRKKAIEV GTRVDALHVI TTTTTTITAS TTMSVAADTV VRETRENIRL RYPAVRQGGA 180 VAGAFGSEGF RENNKVGKNP DLNQRPSPDC SDEEDAKL* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5wx9_A | 2e-16 | 36 | 102 | 8 | 75 | Ethylene-responsive transcription factor ERF096 |
| Search in ModeBase | ||||||
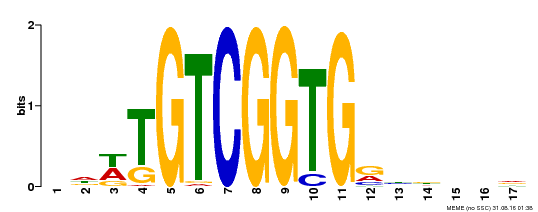
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00276 | DAP | Transfer from AT2G23340 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| TrEMBL | A0A0K9PHY5 | 1e-160 | A0A0K9PHY5_ZOSMR; Ethylene-responsive transcription factor | ||||
| STRING | Traes_2BL_1E57B73B21.5 | 7e-49 | (Triticum aestivum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3943 | 30 | 75 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G06746.1 | 4e-37 | related to AP2 9 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Zosma237g00270.1 |



