 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zosma203g00250.1 | ||||||||
| Common Name | ZOSMA_203G00250 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Alismatales; Zosteraceae; Zostera
|
||||||||
| Family | SRS | ||||||||
| Protein Properties | Length: 376aa MW: 39252.3 Da PI: 8.4257 | ||||||||
| Description | SRS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | DUF702 | 215.2 | 1.5e-66 | 132 | 298 | 3 | 154 |
DUF702 3 sgtasCqdCGnqakkdCaheRCRtCCksrgfdCathvkstWvpaakrrerqqqlaaasskaaasa...............aeaaskrkrelks 80
g++sCqdCGnqakkdC h+RCRtCCksr++ C thvkstWvpaakrrerq+ql ++++ +++++ + +askr+re ++
Zosma203g00250.1 132 GGGMSCQDCGNQAKKDCIHMRCRTCCKSRNLPCPTHVKSTWVPAAKRRERQHQLMSSMQMQQQHHssprllrscggggsgSGEASKRPREITT 224
5789*************************************************99988877777779**********99989999*****999 PP
DUF702 81 kkqsalsstklssaeskkeletsslPeevsseavfrcvrvssvddgeeelaYqtavsigGhvfkGiLydqGlee 154
+ + l++t +++++s + + ++P+evss+a+frcv+vs++dd++e++aYqtav+igGh+fkGiLydqG+++
Zosma203g00250.1 225 LACTRLANTAVTTSNSGGMEIVGTFPSEVSSPAIFRCVKVSAMDDADEQYAYQTAVNIGGHTFKGILYDQGTDS 298
999999999999999998888899***********************************************997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF05142 | 6.9E-64 | 134 | 297 | IPR007818 | Protein of unknown function DUF702 |
| TIGRFAMs | TIGR01623 | 4.8E-26 | 136 | 178 | IPR006510 | Zinc finger, lateral root primordium type 1 |
| TIGRFAMs | TIGR01624 | 2.6E-26 | 249 | 295 | IPR006511 | Lateral Root Primordium type 1, C-terminal |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009938 | Biological Process | negative regulation of gibberellic acid mediated signaling pathway | ||||
| GO:0010051 | Biological Process | xylem and phloem pattern formation | ||||
| GO:0010252 | Biological Process | auxin homeostasis | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048479 | Biological Process | style development | ||||
| GO:0048480 | Biological Process | stigma development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0046982 | Molecular Function | protein heterodimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 376 aa Download sequence Send to blast |
MAGFSLGGGN NNSSSRSNQQ QQPPAEDQIP PENLFFYHSA AAGSARNEEI TTTYAKGFEL 60 WQQHQHHTTQ QHLHHQQQQR HHQFTLYSSG PPGGLGIFGD DPLGGVGIGP SSIVVGSGSG 120 GSSSGRGMRG TGGGMSCQDC GNQAKKDCIH MRCRTCCKSR NLPCPTHVKS TWVPAAKRRE 180 RQHQLMSSMQ MQQQHHSSPR LLRSCGGGGS GSGEASKRPR EITTLACTRL ANTAVTTSNS 240 GGMEIVGTFP SEVSSPAIFR CVKVSAMDDA DEQYAYQTAV NIGGHTFKGI LYDQGTDSQY 300 SGVGGGGHRG DTSSSSPAAI GSSSTAVASA STSAAAAAAA AAASAVPTLM DPSSYYPTPL 360 NSFMSGTQFF PHPRP* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00613 | PBM | Transfer from AT3G51060 | Download |
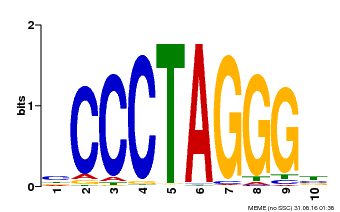
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010271515.1 | 5e-94 | PREDICTED: protein SHI RELATED SEQUENCE 1-like | ||||
| TrEMBL | A0A0K9PLG5 | 0.0 | A0A0K9PLG5_ZOSMR; Lateral root primordium (LRP) protein-related | ||||
| STRING | XP_010271515.1 | 2e-93 | (Nelumbo nucifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1354 | 35 | 124 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G66350.1 | 2e-57 | Lateral root primordium (LRP) protein-related | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Zosma203g00250.1 |



