 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zosma193g00190.1 | ||||||||
| Common Name | ZOSMA_193G00190 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Alismatales; Zosteraceae; Zostera
|
||||||||
| Family | C3H | ||||||||
| Protein Properties | Length: 667aa MW: 73391.5 Da PI: 7.5288 | ||||||||
| Description | C3H family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-CCCH | 24 | 7e-08 | 262 | 282 | 5 | 26 |
--SGGGGTS--TTTTT-SS-SS CS
zf-CCCH 5 lCrffartGtCkyGdrCkFaHg 26
+C+ f++ G C+ Gd C +aHg
Zosma193g00190.1 262 PCPEFRK-GSCRNGDACEYAHG 282
8******.*************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF48403 | 8.71E-13 | 50 | 134 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 2.1E-12 | 50 | 136 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 15.724 | 52 | 135 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 3.2 | 52 | 85 | IPR002110 | Ankyrin repeat |
| Pfam | PF12796 | 1.0E-6 | 52 | 124 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 5.11E-13 | 54 | 135 | No hit | No description |
| PROSITE profile | PS50088 | 10.072 | 92 | 127 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.03 | 92 | 124 | IPR002110 | Ankyrin repeat |
| SMART | SM00356 | 6.3E-5 | 257 | 283 | IPR000571 | Zinc finger, CCCH-type |
| Gene3D | G3DSA:4.10.1000.10 | 1.2E-4 | 261 | 284 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 13.056 | 262 | 284 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 2.1E-5 | 262 | 282 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 6.653 | 292 | 316 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 36 | 292 | 315 | IPR000571 | Zinc finger, CCCH-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 667 aa Download sequence Send to blast |
MESPTTLDPT LLLEYAASDD LVSFRKAVEM DFRTIDVASK WYGRWSSQMG FQERTPLMVA 60 ALYGSVSVLE YILSFETTFP GTIHVNRRCG SDNSSALHLA VSGASPVYFK IVKLLLDAKA 120 DVSLLDSSGR TPIHVVPRQK PCLGKMKSAV ELALGGGHGY FDGLEGEIPT STPSLVNPKQ 180 QKVTSPGAGG LGLGEKKEYP VDATLPDIKS GVCGTDEFRM YTFKVKPCSR AYSHDWTECP 240 FVHPGENARR RDPRKYHYSC VPCPEFRKGS CRNGDACEYA HGVFESWLHP AQYRTRLCKD 300 ETGCNRRVCF FAHKTEELRP LYNSVVSSPR SPANLPSLDL SSAAAMLMNQ QTPLSPSSSS 360 ALAAAVWLNQ NTGTPTSQLP SSRLKSAMNA RDMDYEFDLL GLDGSAKKQV QNQNFIDEIG 420 NLSSRSMNMD YGVDFFGSPE SNPLSPLTNH SLKQSPSLSQ FQSPPSSIQK QLLSNYNNGG 480 GNIPSSPSTR SSSPSFGLDH KLASAIINSR ASSFAKRSQS FVDRGVGNRQ VIGSTQSQAN 540 SDWGSPSGKL DWGIQGEELN KLRKSASFAF RKNEKAIPAS QLDDEPDVSW VQSIVKDGPM 600 FENGYVEDER EKMQRNQQQM MMLMMMNKKK QFHLNSGAGE FHPGDSFASW NPDQIQIHLD 660 QEQMVA* |
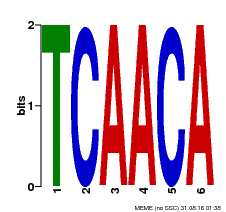
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00566 | DAP | Transfer from AT5G58620 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | JN944375 | 1e-138 | JN944375.1 Zostera marina microsatellite Pzm9 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010937855.1 | 0.0 | zinc finger CCCH domain-containing protein 33 | ||||
| TrEMBL | A0A0K9PNZ2 | 0.0 | A0A0K9PNZ2_ZOSMR; Zinc finger CCCH domain-containing protein | ||||
| STRING | XP_006488539.1 | 0.0 | (Citrus sinensis) | ||||
| STRING | XP_006425080.1 | 0.0 | (Citrus clementina) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP823 | 37 | 126 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G58620.1 | 1e-145 | C3H family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Zosma193g00190.1 |



