 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zosma148g00340.1 | ||||||||
| Common Name | ZOSMA_148G00340 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Alismatales; Zosteraceae; Zostera
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 467aa MW: 50725.9 Da PI: 6.5351 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 86.3 | 3.1e-27 | 189 | 242 | 2 | 56 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
+++WtpeLH+rFveav+qL G ekA+P++ile+m++++Lt+++++SHLQkYR++
Zosma148g00340.1 189 IKVDWTPELHRRFVEAVDQL-GVEKAVPSRILEIMGIDCLTRHNIASHLQKYRSH 242
5899****************.********************************86 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 17.779 | 185 | 244 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 7.0E-26 | 186 | 245 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 2.51E-17 | 187 | 245 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.4E-25 | 188 | 242 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 8.9E-7 | 191 | 240 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007165 | Biological Process | signal transduction | ||||
| GO:0009658 | Biological Process | chloroplast organization | ||||
| GO:0009910 | Biological Process | negative regulation of flower development | ||||
| GO:0010380 | Biological Process | regulation of chlorophyll biosynthetic process | ||||
| GO:0010638 | Biological Process | positive regulation of organelle organization | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 467 aa Download sequence Send to blast |
MLAISPLETA GCNLDGRGVG DAAAEDYVAG GGLVGGGAGN GGGEHDQHHQ HQHQHDSFAD 60 AFSSGNLQLE DINFDDLFVG FDDGTGDILP DLELDPDELL GVFNTSFEED MEMGSLGEVK 120 SGGTESSGGV DGEKEEGDSF KTTSDGGDQT VPETTGEDVK VSESVAAVDE RNRKVRKSTA 180 GGMQKKRKIK VDWTPELHRR FVEAVDQLGV EKAVPSRILE IMGIDCLTRH NIASHLQKYR 240 SHRKHVLARE AEAASWTHRR KIYSPTTSTV GHVVPVLKRE TITPWMGAPA TMGFPTTPLR 300 TQPPPPASPS PSIQSFRPLH VWGHPTVEHH MIHMWPKHVV PSHPPHHHWA PPPHDPSAYW 360 LPPHYQPKTP HYPHHPPLPP QRFSSPSPIA TGIPPFSPPP PAKLHHQFQP TKEIIDKVIG 420 DVLTKPWLPL ALGLKPPSVD SVLVALQREG ITKIPSHPSS SSSLCS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 8e-17 | 186 | 246 | 3 | 63 | ARR10-B |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00022 | PBM | Transfer from AT2G20570 | Download |
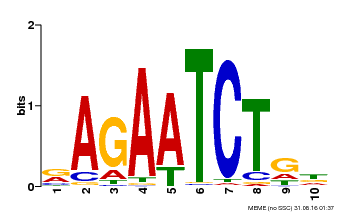
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010260326.1 | 1e-100 | PREDICTED: probable transcription factor GLK1 | ||||
| TrEMBL | A0A0K9PZ41 | 0.0 | A0A0K9PZ41_ZOSMR; Response regulator | ||||
| STRING | XP_010260326.1 | 1e-99 | (Nelumbo nucifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2415 | 38 | 86 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G20570.1 | 7e-59 | GBF's pro-rich region-interacting factor 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Zosma148g00340.1 |



