 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zosma137g00220.1 | ||||||||
| Common Name | ZOSMA_137G00220 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Alismatales; Zosteraceae; Zostera
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 384aa MW: 43522.7 Da PI: 9.6444 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 47 | 5.7e-15 | 39 | 83 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r rWT++E++++++a+ ++G W+ I++++g ++t q++s+ qky
Zosma137g00220.1 39 RERWTADEHQKFIEAISLYGRA-WRCISEYIG-TKTTIQIRSHAQKY 83
78******************88.*********.*************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 4.71E-15 | 34 | 88 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 19.234 | 34 | 88 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 1.6E-15 | 37 | 86 | IPR006447 | Myb domain, plants |
| Gene3D | G3DSA:1.10.10.60 | 6.5E-8 | 38 | 79 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 6.6E-11 | 38 | 86 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.2E-12 | 39 | 83 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.73E-8 | 41 | 84 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 384 aa Download sequence Send to blast |
MCSDPFIMPQ KQLPVETRLS SETENILKVR KPYIITKQRE RWTADEHQKF IEAISLYGRA 60 WRCISEYIGT KTTIQIRSHA QKYFSKIANK SSDSNVDSTV LVEIPPPRQK RKPNHPYPRK 120 FGNASDLKSH VLKKNLEKTT SDQESKTPKS VLSEFSSDFI GSSVSDIQKH KNALSDTCSG 180 VLQPSLAEEN GCTSSQSRDA VEESNPTSRK RRIQLFGTTI YTTEEDSVRG SKTKDEQIPT 240 PSEKGNVTPR VSDGNSPKDS WNPMLYYCMP PQQQDALQFH QNIVPLMPFY GGGFQHHPPA 300 SYQKYDASNE ERSHNKEIQE GSTSSFSYAS PMQTEIKVNH HNSMECHLAQ GKQRLFSSQS 360 QKSISNRQQR LKGFVPYKRP NIS* |
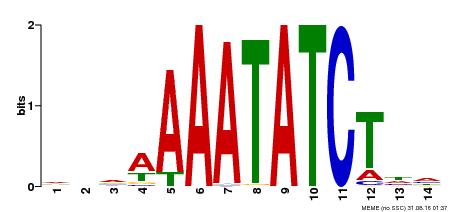
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00515 | DAP | Transfer from AT5G17300 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| TrEMBL | A0A0K9Q0H6 | 0.0 | A0A0K9Q0H6_ZOSMR; Uncharacterized protein | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP7783 | 36 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G17300.1 | 3e-37 | MYB_related family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Zosma137g00220.1 |



