| Signature Domain? help Back to Top |
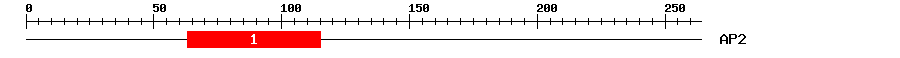 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | AP2 | 49 | 1.5e-15 | 63 | 115 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpseng.krkrfslgkfgtaeeAakaaiaarkkleg 55
+ ++GVr++ grWv+e+r+p g + +r +lg++ ae Aa+a++aa+++l g
GRMZM2G124037_P01 63 PVFRGVRRRGAAGRWVCEVRVP---GrRGARLWLGTYLAAEAAARAHDAAMLALQG 115
579******************9...6455***********************9987 PP
|
| Publications
? help Back to Top |
- Kikuchi S, et al.
Collection, mapping, and annotation of over 28,000 cDNA clones from japonica rice.
Science, 2003. 301(5631): p. 376-9
[PMID:12869764] - Oh SJ,Kwon CW,Choi DW,Song SI,Kim JK
Expression of barley HvCBF4 enhances tolerance to abiotic stress in transgenic rice.
Plant Biotechnol. J., 2007. 5(5): p. 646-56
[PMID:17614953] - Xiao BZ, et al.
Evaluation of seven function-known candidate genes for their effects on improving drought resistance of transgenic rice under field conditions.
Mol Plant, 2009. 2(1): p. 73-83
[PMID:19529831] - Mito T,Seki M,Shinozaki K,Ohme-Takagi M,Matsui K
Generation of chimeric repressors that confer salt tolerance in Arabidopsis and rice.
Plant Biotechnol. J., 2011. 9(7): p. 736-46
[PMID:21114612] - Sun H, et al.
ENAC1, a NAC transcription factor, is an early and transient response regulator induced by abiotic stress in rice (Oryza sativa L.).
Mol. Biotechnol., 2012. 52(2): p. 101-10
[PMID:22161313] - Huang J, et al.
A TFIIIA-type zinc finger protein confers multiple abiotic stress tolerances in transgenic rice (Oryza sativa L.).
Plant Mol. Biol., 2012. 80(3): p. 337-50
[PMID:22930448] - Lourenço T, et al.
Isolation and characterization of rice (Oryza sativa L.) E3-ubiquitin ligase OsHOS1 gene in the modulation of cold stress response.
Plant Mol. Biol., 2013. 83(4-5): p. 351-63
[PMID:23780733] - Chen X, et al.
The NAC family transcription factor OsNAP confers abiotic stress response through the ABA pathway.
Plant Cell Physiol., 2014. 55(3): p. 604-19
[PMID:24399239] - Wang ST, et al.
MicroRNA319 positively regulates cold tolerance by targeting OsPCF6 and OsTCP21 in rice (Oryza sativa L.).
PLoS ONE, 2014. 9(3): p. e91357
[PMID:24667308] - Chen M,Zhao Y,Zhuo C,Lu S,Guo Z
Overexpression of a NF-YC transcription factor from bermudagrass confers tolerance to drought and salinity in transgenic rice.
Plant Biotechnol. J., 2015. 13(4): p. 482-91
[PMID:25283804] - Paul S,Gayen D,Datta SK,Datta K
Dissecting root proteome of transgenic rice cultivars unravels metabolic alterations and accumulation of novel stress responsive proteins under drought stress.
Plant Sci., 2015. 234: p. 133-43
[PMID:25804816] - Liu J, et al.
Functional Analysis of the Maize C-Repeat/DRE Motif-Binding Transcription Factor CBF3 Promoter in Response to Abiotic Stress.
Int J Mol Sci, 2015. 16(6): p. 12131-46
[PMID:26030672] - Challam C,Ghosh T,Rai M,Tyagi W
Allele mining across DREB1A and DREB1B in diverse rice genotypes suggest a highly conserved pathway inducible by low temperature.
J. Genet., 2015. 94(2): p. 231-8
[PMID:26174670] - Kan CC,Chung TY,Juo YA,Hsieh MH
Glutamine rapidly induces the expression of key transcription factor genes involved in nitrogen and stress responses in rice roots.
BMC Genomics, 2015. 16(1): p. 731
[PMID:26407850] - Min HJ,Jung YJ,Kang BG,Kim WT
CaPUB1, a Hot Pepper U-box E3 Ubiquitin Ligase, Confers Enhanced Cold Stress Tolerance and Decreased Drought Stress Tolerance in Transgenic Rice (Oryza sativa L.).
Mol. Cells, 2016. 39(3): p. 250-7
[PMID:26674966] - Kakar KU, et al.
A consortium of rhizobacterial strains and biochemical growth elicitors improve cold and drought stress tolerance in rice (Oryza sativa L.).
Plant Biol (Stuttg), 2016. 18(3): p. 471-83
[PMID:26681628] - Huo C, et al.
Comparative Study of Early Cold-Regulated Proteins by Two-Dimensional Difference Gel Electrophoresis Reveals a Key Role for Phospholipase Dα1 in Mediating Cold Acclimation Signaling Pathway in Rice.
Mol. Cell Proteomics, 2016. 15(4): p. 1397-411
[PMID:26747563] - Dou M,Cheng S,Zhao B,Xuan Y,Shao M
The Indeterminate Domain Protein ROC1 Regulates Chilling Tolerance via Activation of DREB1B/CBF1 in Rice.
Int J Mol Sci, 2016. 17(3): p. 233
[PMID:26927068] - Kudo M, et al.
Double overexpression of DREB and PIF transcription factors improves drought stress tolerance and cell elongation in transgenic plants.
Plant Biotechnol. J., 2017. 15(4): p. 458-471
[PMID:27683092] - He X, et al.
A rice jacalin-related mannose-binding lectin gene, OsJRL, enhances Escherichia coli viability under high salinity stress and improves salinity tolerance of rice.
Plant Biol (Stuttg), 2017. 19(2): p. 257-267
[PMID:27718311]
|




