| Signature Domain? help Back to Top |
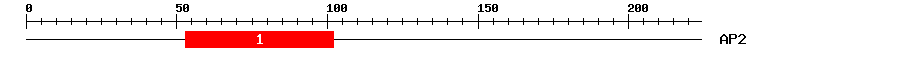 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | AP2 | 57.5 | 3.4e-18 | 53 | 102 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
y+GVr++ +g+Wv+e+r+p +k+ r++lg+f t e+Aa+a++ a+++l+g
AT5G51990.1 53 IYRGVRQRN-SGKWVCEVREP---NKKSRIWLGTFPTVEMAARAHDVAALALRG 102
69****999.7******9998...347*************************98 PP
|
| Annotation --
Nucleotide ? help
Back to Top |
| Source |
Hit ID |
E-value |
Description |
| GenBank | EF523169 | 0.0 | EF523169.1 Arabidopsis thaliana ecotype Oy-0 C-repeat binding factor 4 (CBF4) mRNA, complete cds. |
| GenBank | EF523170 | 0.0 | EF523170.1 Arabidopsis thaliana ecotype Yo-0 C-repeat binding factor 4 (CBF4) mRNA, complete cds. |
| GenBank | EF523171 | 0.0 | EF523171.1 Arabidopsis thaliana ecotype Ag-0 C-repeat binding factor 4 (CBF4) mRNA, complete cds. |
| GenBank | EF523172 | 0.0 | EF523172.1 Arabidopsis thaliana ecotype Ak-1 C-repeat binding factor 4 (CBF4) mRNA, complete cds. |
| GenBank | EF523174 | 0.0 | EF523174.1 Arabidopsis thaliana ecotype Bs-1 C-repeat binding factor 4 (CBF4) mRNA, complete cds. |
| GenBank | EF523175 | 0.0 | EF523175.1 Arabidopsis thaliana ecotype Gre-0 C-repeat binding factor 4 (CBF4) mRNA, complete cds. |
| GenBank | EF523176 | 0.0 | EF523176.1 Arabidopsis thaliana ecotype Co-1 C-repeat binding factor 4 (CBF4) mRNA, complete cds. |
| GenBank | EF523177 | 0.0 | EF523177.1 Arabidopsis thaliana ecotype Can-0 C-repeat binding factor 4 (CBF4) mRNA, complete cds. |
| GenBank | EF523178 | 0.0 | EF523178.1 Arabidopsis thaliana ecotype Cvi-0 C-repeat binding factor 4 (CBF4) mRNA, complete cds. |
| GenBank | EF523179 | 0.0 | EF523179.1 Arabidopsis thaliana ecotype Kin-0 C-repeat binding factor 4 (CBF4) mRNA, complete cds. |
| GenBank | EF523180 | 0.0 | EF523180.1 Arabidopsis thaliana ecotype Lip-0 C-repeat binding factor 4 (CBF4) mRNA, complete cds. |
| GenBank | EF523181 | 0.0 | EF523181.1 Arabidopsis thaliana ecotype Pog-0 C-repeat binding factor 4 (CBF4) mRNA, complete cds. |
| GenBank | EF523182 | 0.0 | EF523182.1 Arabidopsis thaliana ecotype Van-0 C-repeat binding factor 4 (CBF4) mRNA, complete cds. |
| GenBank | EF523183 | 0.0 | EF523183.1 Arabidopsis thaliana ecotype Bla-1 C-repeat binding factor 4 (CBF4) mRNA, complete cds. |
| GenBank | EF523185 | 0.0 | EF523185.1 Arabidopsis thaliana ecotype Hau-0 C-repeat binding factor 4 (CBF4) mRNA, complete cds. |
| GenBank | EF523186 | 0.0 | EF523186.1 Arabidopsis thaliana ecotype Mv-0 C-repeat binding factor 4 (CBF4) mRNA, complete cds. |
| GenBank | EF523188 | 0.0 | EF523188.1 Arabidopsis thaliana ecotype Bur-0 C-repeat binding factor 4 (CBF4) mRNA, complete cds. |
| GenBank | EF523189 | 0.0 | EF523189.1 Arabidopsis thaliana ecotype Per-3 C-repeat binding factor 4 (CBF4) mRNA, complete cds. |
| GenBank | EF523191 | 0.0 | EF523191.1 Arabidopsis thaliana ecotype Sapporo-0 C-repeat binding factor 4 (CBF4) mRNA, complete cds. |
| GenBank | EF523192 | 0.0 | EF523192.1 Arabidopsis thaliana ecotype Tscha-1 C-repeat binding factor 4 (CBF4) mRNA, complete cds. |
| Publications
? help Back to Top |
- Riechmann JL, et al.
Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes.
Science, 2000. 290(5499): p. 2105-10
[PMID:11118137] - Sakuma Y, et al.
DNA-binding specificity of the ERF/AP2 domain of Arabidopsis DREBs, transcription factors involved in dehydration- and cold-inducible gene expression.
Biochem. Biophys. Res. Commun., 2002. 290(3): p. 998-1009
[PMID:11798174] - Hanano S, et al.
Analysis of gene expression in Arabidopsis thaliana by array hybridization with genomic DNA fragments aligned along chromosomal regions.
Plant J., 2002. 30(2): p. 247-55
[PMID:12000460] - Haake V, et al.
Transcription factor CBF4 is a regulator of drought adaptation in Arabidopsis.
Plant Physiol., 2002. 130(2): p. 639-48
[PMID:12376631] - Vandepoele K,Simillion C,Van de Peer Y
Detecting the undetectable: uncovering duplicated segments in Arabidopsis by comparison with rice.
Trends Genet., 2002. 18(12): p. 606-8
[PMID:12446138] - Dal Bosco C, et al.
Inactivation of the chloroplast ATP synthase gamma subunit results in high non-photochemical fluorescence quenching and altered nuclear gene expression in Arabidopsis thaliana.
J. Biol. Chem., 2004. 279(2): p. 1060-9
[PMID:14576160] - Navarro L, et al.
The transcriptional innate immune response to flg22. Interplay and overlap with Avr gene-dependent defense responses and bacterial pathogenesis.
Plant Physiol., 2004. 135(2): p. 1113-28
[PMID:15181213] - Lee BH,Henderson DA,Zhu JK
The Arabidopsis cold-responsive transcriptome and its regulation by ICE1.
Plant Cell, 2005. 17(11): p. 3155-75
[PMID:16214899] - Suzuki N, et al.
Enhanced tolerance to environmental stress in transgenic plants expressing the transcriptional coactivator multiprotein bridging factor 1c.
Plant Physiol., 2005. 139(3): p. 1313-22
[PMID:16244138] - Nakano T,Suzuki K,Fujimura T,Shinshi H
Genome-wide analysis of the ERF gene family in Arabidopsis and rice.
Plant Physiol., 2006. 140(2): p. 411-32
[PMID:16407444] - Xiong Y,Fei SZ
Functional and phylogenetic analysis of a DREB/CBF-like gene in perennial ryegrass (Lolium perenne L.).
Planta, 2006. 224(4): p. 878-88
[PMID:16614820] - Underwood BA,Vanderhaeghen R,Whitford R,Town CD,Hilson P
Simultaneous high-throughput recombinational cloning of open reading frames in closed and open configurations.
Plant Biotechnol. J., 2006. 4(3): p. 317-24
[PMID:17147637] - Zhou N,Robinson SJ,Huebert T,Bate NJ,Parkin IA
Comparative genome organization reveals a single copy of CBF in the freezing tolerant crucifer Thlaspi arvense.
Plant Mol. Biol., 2007. 65(5): p. 693-705
[PMID:17899397] - Jung C, et al.
Overexpression of AtMYB44 enhances stomatal closure to confer abiotic stress tolerance in transgenic Arabidopsis.
Plant Physiol., 2008. 146(2): p. 623-35
[PMID:18162593] - Welling A,Palva ET
Involvement of CBF transcription factors in winter hardiness in birch.
Plant Physiol., 2008. 147(3): p. 1199-211
[PMID:18467468] - Lin YH, et al.
Molecular population genetics and gene expression analysis of duplicated CBF genes of Arabidopsis thaliana.
BMC Plant Biol., 2008. 8: p. 111
[PMID:18990244] - Wang Y,Hua J
A moderate decrease in temperature induces COR15a expression through the CBF signaling cascade and enhances freezing tolerance.
Plant J., 2009. 60(2): p. 340-9
[PMID:19563440] - Abdeen A,Schnell J,Miki B
Transcriptome analysis reveals absence of unintended effects in drought-tolerant transgenic plants overexpressing the transcription factor ABF3.
BMC Genomics, 2010. 11: p. 69
[PMID:20105335] - Causier B,Ashworth M,Guo W,Davies B
The TOPLESS interactome: a framework for gene repression in Arabidopsis.
Plant Physiol., 2012. 158(1): p. 423-38
[PMID:22065421] - Nakai Y, et al.
Vascular plant one-zinc-finger protein 1/2 transcription factors regulate abiotic and biotic stress responses in Arabidopsis.
Plant J., 2013. 73(5): p. 761-75
[PMID:23167462] - Efroni I, et al.
Regulation of leaf maturation by chromatin-mediated modulation of cytokinin responses.
Dev. Cell, 2013. 24(4): p. 438-45
[PMID:23449474] - Ding Y, et al.
Four distinct types of dehydration stress memory genes in Arabidopsis thaliana.
BMC Plant Biol., 2013. 13: p. 229
[PMID:24377444] - Guttikonda SK, et al.
Overexpression of AtDREB1D transcription factor improves drought tolerance in soybean.
Mol. Biol. Rep., 2014. 41(12): p. 7995-8008
[PMID:25192890] - Jin J, et al.
An Arabidopsis Transcriptional Regulatory Map Reveals Distinct Functional and Evolutionary Features of Novel Transcription Factors.
Mol. Biol. Evol., 2015. 32(7): p. 1767-73
[PMID:25750178] - Zhang T, et al.
Overexpression of a NF-YB3 transcription factor from Picea wilsonii confers tolerance to salinity and drought stress in transformed Arabidopsis thaliana.
Plant Physiol. Biochem., 2015. 94: p. 153-64
[PMID:26093308] - Carlow CE, et al.
Nuclear localization and transactivation by Vitis CBF transcription factors are regulated by combinations of conserved amino acid domains.
Plant Physiol. Biochem., 2017. 118: p. 306-319
[PMID:28675818] - Li B, et al.
Network-Guided Discovery of Extensive Epistasis between Transcription Factors Involved in Aliphatic Glucosinolate Biosynthesis.
Plant Cell, 2018. 30(1): p. 178-195
[PMID:29317470]
|





